Abstract
The ongoing coronavirus disease 2019 (COVID-19) pandemic, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection, poses a severe threat to humanity. Rapid and comprehensive analysis of both pathogen and host sequencing data is critical to track infection and inform therapies. In this study, we performed unbiased metatranscriptomic analysis of clinical samples from COVID-19 patients using a recently developed RNA-seq library construction method (TRACE-seq), which utilizes tagmentation activity of Tn5 on RNA/DNA hybrids. This approach avoids the laborious and time-consuming steps in traditional RNA-seq procedure, and hence is fast, sensitive, and convenient. We demonstrated that TRACE-seq allowed integrated characterization of full genome information of SARS-CoV-2, putative pathogens causing coinfection, antibiotic resistance, and host response from single throat swabs. We believe that the integrated information will deepen our understanding of pathogenesis and improve diagnostic accuracy for infectious diseases.
Similar content being viewed by others
Introduction
Longstanding, emerging, and re-emerging infectious diseases continuously threaten human health across centuries1. Precise and rapid identification of pathogens from clinical samples is important for both guiding infection treatment strategies and monitoring novel infectious disease outbreaks, e.g., the outbreak of SARS-CoV-2, in the community. While most nucleic acid amplification-based and pathogen-specific antibody detection-based molecular techniques only detect a limited number of pathogens and need their prior knowledge, metagenomic or metatranscriptomic approaches allow for comprehensive and unbiased identification and characterization of microbiome directly from clinical specimens2.
Compared to metagenomic sequencing, metatranscriptomic sequencing has several distinct advantages: it permits detection of RNA viruses that would not be interpreted in metagenomic data, reveals transcriptionally active organism(s) which are more etiologically important, and indicates host immune response which is essential to distinguish true pathogens from colonizers3,4,5. However, the laborious and time-consuming steps in traditional RNA sequencing (RNA-seq) experiments hinder the development of metatranscriptomics-based clinical diagnostics for rapid pathogen identification.
Very recently, we and others have independently developed a rapid and cost-effective RNA-seq method, based on Tn5 tagmentation activity towards RNA/DNA hybrids6,7. Our method, termed âTRACE-seqâ, enables rapid one-tube library construction for RNA-seq experiments and shows excellent performance in comparison to traditional RNA-seq methods. We thus envisioned that this convenient and sensitive method could be applied to clinical specimens for unbiased metatranscriptomic analysis. In this study, we modified the TRACE-seq procedure, shortened the total time, and optimized analytical pipeline to meet the needs for clinical metatranscriptomic diagnosis and analysis. We then applied TRACE-seq to metatranscriptomic sequencing of single throat swab specimens from COVID-19 patients and healthy individuals. We found library construction of specimens could be accomplished in ~2âh with high quality. Analysis of TRACE-seq metatranscriptomic data of 13 SARS-CoV-2-positive samples and 2 negative samples demonstrated the success of this method to sensitively detect SARS-CoV-2 with high coverage even for samples with low virus abundance, or to assemble unknown microbe genome de novo (using SARS-CoV-2 as an example). Moreover, TRACE-seq sensitively detected the microbiome and simultaneously allowed for interrogating antibiotic resistance and host responses. Taken together, TRACE-seq enables unbiased pathogen detection and could have broad applications in the metatranscriptomic study and clinical diagnosis.
Results
TRACE-seq enables metatranscriptomic analysis
To perform metatranscriptomic analysis on clinical samples, such as throat swabs in this study, we made several modifications to TRACE-seq. First, to achieve unbiased sequencing of microbiome, we used both random hexamer and oligo d(T)23VN primers for reverse transcription, using approximately 1/10 total RNA extracted from a single throat swab as input. Secondly, we reduced the total time of library construction to around 2âh (Fig. 1a), which enables TRACE-seq to be more compatible for clinical use, especially when substantial numbers of specimens require investigation. Third, we developed a tailored analytical pipeline of TRACE-seq to simultaneously identify known and unknown pathogens and at the meanwhile to characterize host transcriptional response in a single metatranscriptomic profiling reaction (Fig. 1b). This new pipeline allowed us to obtain rich information from the metatranscriptomic data generated by the modified TRACE-seq.
Sensitive detection of SARS-CoV-2 genome
Since the throat swab samples were from patients with confirmed or suspected COVID-19, we asked whether the untargeted metatranscriptomic sequencing could yield a full genome sequence of SARS-CoV-2 virus. After removing low-quality reads and human reads, the remaining reads were mapped to the SARS-CoV-2 reference genome Wuhan-Hu-1 (accession number: NC_045512). Sequencing covered the reference genome from 7134âbp to 29,903âbp (23.86%â100%), with a genome-wide average sequencing depth from 0.69Ã to 129,901Ã (Supplementary Table S1). Subsequent correlation analysis revealed that the proportion of obtained reads of SARS-CoV-2, the coverage to the reference genome, the average sequencing depth and the median sequencing depth all showed a significant negative correlation with the cycle threshold (Ct) value of SARS-CoV-2 in all samples (Spearman test, Pâ<â0.01) (Fig. 2a). In addition, the nearly whole genome sequence (> 99%) could be acquired from mapping-based approach for most samples with the Ct value below 35 (nâ=â12, 85.7% of 14 samples with the Ct value below 35), with the average sequencing depth of 16,900Ã (from 28.3Ã to 129,901Ã, median depth 1894.96Ã). Moreover, in 7 samples (50%), genomes were completely covered (Fig. 2b; Supplementary Fig. S1 and Table S1).
a Correlation between SARS-CoV-2 sequencing data and Ct values. From the left to the right: the correlation between the ratio of SARS-CoV-2 reads, the coverage of SARS-CoV-2 genome, the average sequencing depth, the median sequencing depth, and the Ct value of each sample. Linear regression indicates the relationship between the sequencing data and the Ct value of samples. b Genome coverage of sequenced samples across the SARS-CoV-2 genome. The x-axis represents the viral genome position, the y-axis represents the log10 depth of each site. Lines in blue represent the median sequencing depth, and areas in gray represent 25th to 75th percentile of sequencing depth. c De novo assembly results of SARS-CoV-2. The x-axis represents each sample, and the y-axis represents log10 lengths of contigs matching SARS-CoV-2. Boxplots represent the length distribution of contigs matching SARS-CoV-2. Dots in different colors represent the number of error bases (shown in legends) in each contig relative to previously known genome sequences.
Reconstruction of full-length genome of SARS-CoV-2
To determine the accuracy of this method in de novo acquisition of pathogen genome, after de novo assembly from non-human reads and BLAST, contigs matching SARS-CoV-2 with a length ⤠100âbp were discarded, and thus 64 contigs were determined to be SARS-CoV-2 genomes or genome fragments. These contigs ranged from 185âbp to 29,835âbp in length, with an average length of 6437âbp. As the Ct value of SARS-CoV-2 increases, SARS-CoV-2 contigs tended to be much more in number and shorter in length. Most of contigs (46, 71.9%) were exactly the same with our previously known viral genomes (Fig. 2c), and some (10, 15.6%) had 1 base difference (mismatch or gap) and the rest had 2â12 base differences. In samples with SARS-CoV-2 Ct value lower than 32, almost full-length genome (29,776â29,835âbp) were obtained just from de novo assembly. Thus, TRACE-seq could enable the de novo assembly of the complete genome of unknown pathogens and be readily utilized to identify emerging pathogens in patients with unknown etiology of infection and efficiently complement routine diagnostics.
Unbiased identification of putative pathogens in addition to SARS-CoV-2
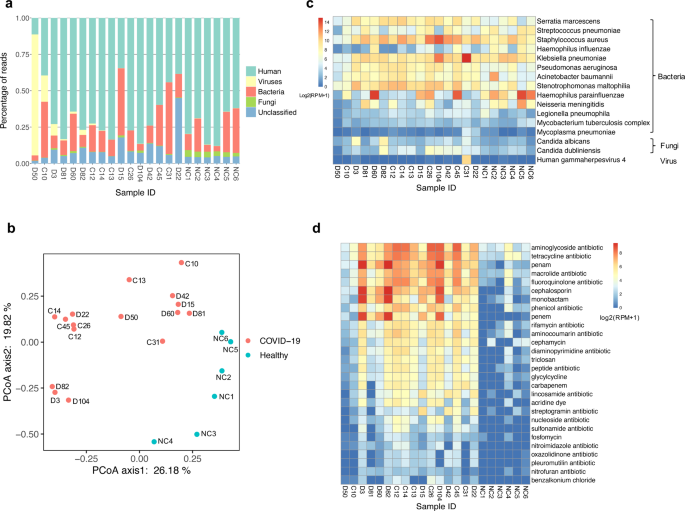
It is widely reported that coinfection (multi-species infection) contributes to enhanced morbidity and mortality, especially in elderly and immunosuppressed influenza patients8,9. Thus, we were curious to see if our metatranscriptomic sequencing approach could capture other pathogens in addition to SARS-CoV-2. Indeed, alignment of TRACE-seq data to microbe reference databases identified many bacteria, fungi, and viruses in both patient and healthy samples (Fig. 3a). To assess whether COVID-19 patients and healthy individuals have different microbe community in their throat, principal coordinates analysis (PCoA) was conducted using relative abundance of the microbiome. We observed that COVID-19 patients harbored a throat microbiome quite different from that of healthy individuals (Fig. 3b). The relative abundance of probable respiratory pathogens was further investigated.
a Histogram showing percentage of reads mapping to human, viruses, bacteria, and fungi for the individual samples. b PCoA of microbiome using relative abundance at the genus level. c Heatmap showing relative abundance of potential respiratory pathogens identified in SARS-CoV-2-positive and -negative samples. RPM, reads per million non-host reads. d Heatmap displaying relative abundance of antibiotic resistance genes in SARS-CoV-2-positive and -negative samples.
Among the probable respiratory pathogens listed in Fig. 3c, Stenotrophomonas maltophilia, Haemophilus parainfluenzae, Staphylococcus aureus, Streptococcus pneumoniae, Haemophilus influenzae, and Acinetobacter baumannii are common commensal organisms of the normal oropharynx; however, they can also become opportunistic pathogens and cause infectious diseases, such as endocarditis, bacteremia, and pneumonia10,11,12,13. Serratia marcescens, Klebsiella pneumoniae, Stenotrophomonas maltophilia, Pseudomonas aeruginosa, Neisseria meningitidis, and Legionella pneumophila cause disease infrequently in normal hosts but can be a major cause of infection in patients with underlying or immunocompromising conditions14,15,16,17,18,19. Mycoplasma pneumoniae is a type of âatypicalâ bacteria that commonly causes mild infections of the respiratory system20. The Mycobacterium tuberculosis complex (MTC or MTBC) is a genetically related group of Mycobacterium species that can cause tuberculosis in humans or other animals21. As for identified fungi, Candida dubliniensis and Candida albicans are both opportunistic yeast and can be detected in the gastrointestinal tract in healthy adults; they were also known to cause respiratory diseases22,23,24. Human gammaherpesvirus 4 is one of the most common viruses in human. It is best known as the cause of infectious mononucleosis25,26, and is also constantly detected in lungs of patients with idiopathic pulmonary fibrosis27. In our results, a relatively high abundance of Serratia marcescens, Staphylococcus aureus, Stenotrophomonas maltophilia, Acinetobacter baumannii, Pseudomonas aeruginosa and Candida dubliniensis were identified in several SARS-CoV-2-positive samples compared with negative samples, which indicated potential coinfection. Nevertheless, these data by themselves could not prove that COVID-19 patients were coinfected by these identified microorganisms; these data have to be carefully interpreted in the clinical context.
Expression profiles of antibiotic resistance genes
Antimicrobial resistance has become a global issue. Pathogens with antibiotic resistance are increasing and many pathogens are becoming multidrug resistant28,29. To characterize antibiotic resistance gene expression profiles, we aligned metatranscriptomic reads against the Comprehensive Antibiotic Resistance Database (CARD)30. On average, transcripts of ~124 antibiotic resistance genes were identified in SARS-CoV-2-positive samples, while only ~15 genes were identified in negative samples. According to the CARD, the identified antibiotic resistance genes confer resistance to 28 classes of antibiotics. Almost all resistance gene classes were more abundant in COVID-19 patients compared to healthy individuals. Genes conferring resistance to beta-lactam (including penam, cephalosporin, monobactam, penem, etc.), aminoglycoside, tetracycline, phenicol, rifamycin, fluoroquinolone, and macrolide were the most abundant (Fig. 3d). Overall, the distinct microbiome, emergence of potential coinfection, and the elevated abundance of antibiotic resistance genes provide new data for establishing clinical therapeutic scheme during the treatment for COVID-19 patients.
Characterization of host response to SARS-CoV-2
Distinguishing infection from colonization remains challenging. Because host transcriptional profiling has emerged as a promising diagnostic tool for infectious diseases31,32, we next tested whether the host response to SARS-CoV-2 could be simultaneously characterized by TRACE-seq-mediated metatranscriptomic analysis from throat swabs. As shown in Fig. 3a, a substantial percentage of the reads are derived from human, and an average of 11,460 human genes with FPKMâ>â1 were detected per sample (Fig. 4a; Supplementary Fig. S2a, b). Based on the host gene expression profiles, the relationships between samples were inspected using a multidimensional scaling (MDS) plot (Fig. 4b). As expected, SARS-CoV-2-positive samples were clearly separated from negative samples. In addition, SARS-CoV-2-positive samples could be divided into two subgroups. Further investigation revealed that the two subgroups could also be separated according to viral load (defined by the Ct value of SARS-CoV-2 ORF1b region target): subgroup 1 with higher viral load (Ct: 21.97â30.25, except sample D15 (Ctâ=â32.5)), and subgroup 2 with lower viral load (Ct: 30.51â37.31). To characterize the host responses to different SARS-CoV-2 viral loads, we performed differential gene expression analysis between low SARS-CoV-2 viral load and negative samples, as well as between high viral load and negative samples. We identified 522 differentially expressed genes between low viral load and negative samples, among which 251 genes were up-regulated in low viral load samples (upper panel, Fig. 4c; Supplementary Fig. S2c). We also identified 402 differentially expressed genes between high viral load and negative samples, among which 225 genes were up-regulated in high viral load samples (lower panel, Fig. 4c; Supplementary Fig. S2d). Gene Ontology (GO) enrichment analysis identified that the top up-regulated biological processes in low viral load samples are cell surface receptor signaling pathway, locomotion, response to external stimulus, defense response and immune response, chemotaxis, movement of cell or subcellular component, localization of cell, positive regulation of NF-kappaB import into nucleus, etc., which indicated that host responses in these patients mainly consist of immune response and recruitment of the immune cells. Nevertheless, the top up-regulated biological processes identified in high viral load samples were defense response, response to external stimulus, biotic stimulus and other organism, immune response, and response to cytokine, etc., which indicated that host responses in these patients were dominated by strong immune response (Fig. 4d; Supplementary Fig. S2d). Further investigation of immune response revealed a subset of up-regulated genes in both low and high viral load samples involved in IL1B-associated inflammatory response (IL1B, CXCR1, CXCR2, FOS, C5AR1, TLR4, CEBPB, MEFV, FPR1, FPR2, SLC11A1, PROK2, PTGS2, OSM, SERPINA1, TNFRSF10C). Besides, several inflammatory response-related genes (CCRL2, NFAM1, FFAR2, AOC3, MMP25, FCER1G, SIGLEC1, TLR2, TLR8, MYLK3) were mainly up-regulated in low viral load samples, and most of them encode proteins functioning as receptors. In addition, several inflammatory response-associated genes (CCL3, NOS2, NUPR1, ALOX5AP) were mainly up-regulated in high viral load samples. Moreover, another subset of genes up-regulated in both low and high viral load samples (ISG15, EGR1, IFIT1, IFIT2, IFIT3, IFITM1, IFITM2, IFITM3, RSAD2) were enriched in type I interferon signaling pathway, with two genes (ISG20, OASL) mainly up-regulated in high viral load samples (Fig. 4e). These results were highly consistent with the previously reported host response to SARS-CoV-233,34,35. Overall, metatranscriptomic data obtained via TRACE-seq of throat swab samples demonstrate reliable characterization of host transcriptional response to the infection of SARS-CoV-2.
a Bar plot showing gene numbers detected in each sample. b MDS plot showing variation among samples based on host transcriptional profiles. c Volcano plots showing differentially expressed genes between low SARS-CoV-2 viral load and negative samples (upper), and between high SARS-CoV-2 viral load and negative samples (lower), respectively. Significantly up- and down-regulated genes (padjâ<â0.05, |log2FoldChange|â> 1) are highlighted in red and blue, respectively. d Bar plots of the most enriched GO terms in low and high SARS-CoV-2 viral load samples, respectively. e Heatmap presenting the significantly up-regulated immune response-related genes in SARS-CoV-2-positive samples compared to negative samples.
Discussion
Although next generation sequencing holds a great potential to directly detect known and unknown pathogens including viruses, bacteria, fungi, and parasites in a single application, the laborious and time-consuming steps in traditional RNA library construction procedure hinders its clinical application. As a rapid and convenient one-tube RNA-seq library construction method, TRACE-seq showed comparable performance as traditional RNA library preparation methods in terms of microbiome and host transcriptome profiling (Supplementary Fig. S3), but significantly lower the barrier for extensive application of unbiased RNA-seq in clinical diagnosis. In addition, multiplexing libraries by utilizing Tn5 transposase containing barcoded adaptors could enable sample investigation in a high-throughput manner, particularly when comprehensive surveillance for emerging pathogens is needed during a sudden disease outbreak.
It is very challenging to discriminate pathogens from background commensal microbiota, since substantive bacteria or fungi can colonize multiple body sites of healthy individuals. The microbe present at a relatively higher abundance in patients compared to healthy individuals is often considered as a pathogen, yet the abundance threshold indicating infection is difficult to define based solely on microbiome information. On the other hand, host transcriptional profiling has been reported to distinguish infectious and noninfectious diseases32 and to further discriminate between virial and bacterial infections31. A previous study integrates host response and unbiased microbe detection for lower respiratory tract infection diagnosis in critically ill adults, using both RNA-seq and DNA-seq but yet lacking antibiotic resistance analysis3. Another study characterized microbial gene expression profiles (including antibiotic resistance genes) using nasal and throat swab samples, and host response using blood samples during influenza infection36. To our knowledge, this is the first study integrating unbiased pathogen detection, antibiotic resistance, and host response analyses in a single approach with throat swabs from COVID-19 patients. In our results, SARS-CoV-2-positive and -negative samples differed significantly in both microbiome composition and host response. Moreover, TRACE-seq hold the potential to construct a network of microbiome composition, antibiotic resistance, and host response for characterizing the complex hostâmicrobiome interactions. Ideally, TRACE-seq data can be utilized to develop a model combining pathogen diversity metric, antibiotic resistance, and host transcriptional classifier for infectious disease diagnosis. We believe that the integrated information acquired from a TRACE-seq library will deepen our understanding of pathogenesis, improve diagnostic accuracy and more precisely inform optimal antimicrobial treatment for infectious diseases caused by not only SARS-CoV-2 but also other pathogens, and eventually facilitate the utility of metatranscriptomic profiling as a routine diagnostic method.
Materials and methods
Ethics statement
The study and use of all samples were approved by the Ethics Committee of Wuhan Institute of Virology (No. WIVH17202001).
Sample collection and nucleotide extraction
Respiratory specimens (swabs) collected from patients admitted to various Wuhan health care facilities were immediately placed into sterile tubes containing 3âmL of viral transport media (VTM). The swabs were deactivated by heating at 56â°C for 30âmin in a biosafety level 2 (BSL 2) laboratory at the Wuhan Institute of Virology in Zhengdian Park with personal protection equipment for biosafety level 3 (BSL 3) laboratory. Total nucleic acids were extracted using QIAamp 96 virus Qiacube HT kit on QIAxtractor Automated extraction (Qiagen, US) following the manufacturerâs instructions.
TRACE-seq library preparation and sequencing
TRACE-seq libraries were constructed using TruePrep® RNA Library Prep Kit for Illumina (Vazyme, TR502-01) according to the manufacturerâs instructions with several modifications. Firstly, 1/10 volume of total nucleic acids extracted from each swab was used for each library without rRNA removal. Secondly, both random hexamers and oligo(dT)20VN primers were used during the reverse transcription process. Thirdly, we used N5 and N7 PCR primer with a final concentration of 0.2âμM during the PCR process. Lastly, after 18 PCR cycles, the library was purified using 0.8à Agencourt AMPure XP beads (Beckman Coulter) and eluted in 20âμL nuclease-free water. The concentration of resulting libraries was determined by Qubit 3.0 fluorometer with the Qubit dsDNA HS Assay kit (Invitrogen) and the size distribution of libraries was assessed by Agilent 2100 Bioanalyzer. Finally, libraries were sequenced on the Illumina Hiseq X10 platform which generated 2 à 150âbp of paired-end raw reads.
NEBNext Ultra II RNA library preparation
NEBNext Ultra II RNA libraries were constructed using NEBNext Ultra II RNA Library Prep Kit for Illumina (NEB, #E7770S) according to the manufacturerâs instructions.
Data preprocessing
Raw reads from sequencing were firstly subjected to Trim Galore (v0.6.4_dev) (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/) for quality control and adaptor trimming. The minimal threshold of quality was 20, and the minimal length of reads to remain was set as 20ânt.
Host transcriptional profiling analysis
Clean reads were firstly mapped to human rRNA sequences using Bowtie2 (v2.2.9)37, and then unmapped reads were mapped to human genome (hg19) and transcriptome using STAR (v2.7.1a)38. The FPKM value for annotated genes was calculated by cuffnorm (v2.2.1)39, and genes with FPKMâ>â1 were considered to be expressed. Multidimensional scaling and differential gene expression analysis were conducted using EdgeR (v3.28.1)40 with gene count data generated by HTSeq (v0.11.2)41. GO enrichment analysis for biological processes was performed by DAVID (v6.8)42 with all significantly up-regulated genes as input. Due to the redundancy of enriched GO terms, GO terms and their P values were further summarized using REViGO43. The top 10 enriched representative GO terms were plotted.
Discrimination and de novo assembly of SARS-CoV-2
After removal of human reads, the remaining data were aligned to the reference genome of Wuhan-Hu-1 (GenBank accession number: NC_045512) using Bowtie2 (v2.2.9)37 for SARS-CoV-2 identification. The coverage and sequencing depth of SARS-CoV-2 genome were calculated by Samtools (v1.9)44. On the other hand, to verify that the method could screen for aetiologic agents and obtain pathogen genome, all non-human reads were processed for de novo assembly using MEGAHIT (v1.2.9) with default parameters45, and then all contigs were searched against NCBI nt database using blastn for classification46. As for accuracy of assembly sequences, contigs determined to come from SARS-CoV-2 were performed blastn (with the parameter â-outfmt 3â) to display the differences with corresponding genome.
Microbiome analysis
After removing human reads, the remaining reads were subjected to microbial taxonomic classification using Kraken2 (v2.0.8-beta)47 with a custom database. To build the custom database, standard RefSeq complete bacterial genomes were downloaded through âkraken2-build --download-library bacteriaâ and complete genomes of human viruses and genome assemblies of fungi were downloaded from NCBIâs RefSeq and added to the custom databaseâs genomic library using the â--add-to-libraryâ switch. PCoA of relative abundances of microbial taxa at the genus level was done using cmdscale command in R. Distances between samples were calculated using Morisita-horn dissimilarity index by vegdist command from vegan package version 2.5â6 (https://CRAN.R-project.org/package=vegan). The antibiotic resistance genes were annotated by aligning the filtered metatranscriptomic reads to the CARD. Antibiotic resistance genes with more than 10 completely matching reads were considered. The relative expression of antibiotic resistance genes was determined as RPM (reads per million non-host reads). All corresponding graphs were plotted using R scripts by RStudio (v1.2.5033) (https://rstudio.com/).
Correlation analysis
The pearson correlation coefficients between TRACE-seq and NEBNext Ultra II RNA kit sequencing data were calculated using function rcorr() in Hmisc package in R, based on microbial relative abundance at genus level called by Kraken2 (v2.0.8-beta)47, and host gene expression profiles generated by cuffnorm (v2.2.1)39.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive48 in National Genomics Data Center49, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession number HRA000650 that are publicly accessible at http://bigd.big.ac.cn/gsa-human.
Change history
10 August 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41421-022-00448-5
References
Lozano, R. et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380, 2095â2128 (2012).
Chiu, C. Y. & Miller, S. A. Clinical metagenomics. Nat. Rev. Genet. 20, 341â355 (2019).
Langelier, C. et al. Integrating host response and unbiased microbe detection for lower respiratory tract infection diagnosis in critically ill adults. Proc. Natl. Acad. Sci. 115, E12353âE12362 (2018).
Langelier, C. et al. Metagenomic sequencing detects respiratory pathogens in hematopoietic cellular transplant patients. Am. J. Respir. Crit. Care Med. 197, 524â528 (2018).
Gliddon, H. D., Herberg, J. A., Levin, M. & Kaforou, M. Genome-wide host RNA signatures of infectious diseases: discovery and clinical translation. Immunology 153, 171â178 (2018).
Lu, B. et al. Transposase-assisted tagmentation of RNA/DNA hybrid duplexes. Elife 9, e54919 (2020).
Di, L. et al. RNA sequencing by direct tagmentation of RNA/DNA hybrids. Proc. Natl. Acad. Sci. USA 117, 2886â2893 (2020).
Fischer, N. et al. Evaluation of unbiased next-generation sequencing of RNA (RNA-seq) as a diagnostic method in influenza virus-positive respiratory samples. J. Clin. Microbiol. 53, 2238â2250 (2015).
Chertow, D. S. & Memoli, M. J. Bacterial coinfection in influenza: a grand rounds review. JAMA 309, 275â282 (2013).
Man, W. H., de Steenhuijsen Piters, W. A. & Bogaert, D. The microbiota of the respiratory tract: gatekeeper to respiratory health. Nat. Rev. Microbiol 15, 259â270 (2017).
Mitchell, J. L. & Hill, S. L. Immune response to Haemophilus parainfluenzae in patients with chronic obstructive lung disease. Clin. Diagn. Lab Immunol. 7, 25â30 (2000).
Smeltzer, M. S. Staphylococcus aureus pathogenesis: the importance of reduced cytotoxicity. Trends Microbiol. 24, 681â682 (2016).
Howard, A., OâDonoghue, M., Feeney, A. & Sleator, R. D. Acinetobacter baumannii: an emerging opportunistic pathogen. Virulence 3, 243â250 (2012).
Paczosa, M. K. & Mecsas, J. Klebsiella pneumoniae: going on the offense with a strong defense. Microbiol. Mol. Biol. Rev. 80, 629â661 (2016).
Brooke, J. S. Stenotrophomonas maltophilia: an emerging global opportunistic pathogen. Clin. Microbiol. Rev. 25, 2â41 (2012).
Sadikot, R. T., Blackwell, T. S., Christman, J. W. & Prince, A. S. Pathogen-host interactions in Pseudomonas aeruginosa pneumonia. Am. J. Respir. Crit. Care Med. 171, 1209â1223 (2005).
Overturf, G. D. Indications for the immunological evaluation of patients with meningitis. Clin. Infect. Dis. 36, 189â194 (2003).
Kumpers, P. et al. Legionnairesâ disease in immunocompromised patients: a case report of Legionella longbeachae pneumonia and review of the literature. J. Med. Microbiol. 57, 384â387 (2008).
Maki, D. G., Hennekens, C. G., Phillips, C. W., Shaw, W. V. & Bennett, J. V. Nosocomial urinary tract infection with Serratia marcescens: an epidemiologic study. J. Infect. Dis. 128, 579â587 (1973).
Kashyap, S. & Sarkar, M. Mycoplasma pneumonia: clinical features and management. Lung India 27, 75â85 (2010).
Chiner-Oms, A. et al. Genome-wide mutational biases fuel transcriptional diversity in the Mycobacterium tuberculosis complex. Nat. Commun. 10, 3994 (2019).
AbdulWahab, A., Salah, H., Chandra, P. & Taj-Aldeen, S. J. Persistence of Candida dubliniensis and lung function in patients with cystic fibrosis. BMC Res. Notes 10, 326 (2017).
Shweihat, Y., Perry, J. 3rd & Shah, D. Isolated Candida infection of the lung. Respir. Med. Case Rep. 16, 18â19 (2015).
Wahab, A. A. et al. High prevalence of Candida dubliniensis in lower respiratory tract secretions from cystic fibrosis patients may be related to increased adherence properties. Int. J. Infect. Dis. 24, 14â19 (2014).
Stanfield, B. A. & Luftig, M. A. Recent advances in understanding Epstein-Barr virus. F1000Res 6, 386 (2017).
Dunmire, S. K., Hogquist, K. A. & Balfour, H. H. Infectious Mononucleosis. Curr. Top. Microbiol. Immunol. 390, 211â240 (2015).
Tang, Y. W. et al. Herpesvirus DNA is consistently detected in lungs of patients with idiopathic pulmonary fibrosis. J. Clin. Microbiol. 41, 2633â2640 (2003).
Boolchandani, M., DâSouza, A. W. & Dantas, G. Sequencing-based methods and resources to study antimicrobial resistance. Nat. Rev. Genet. 20, 356â370 (2019).
Tillotson, G. S. & Zinner, S. H. Burden of antimicrobial resistance in an era of decreasing susceptibility. Expert Rev. Anti Infect. Ther. 15, 663â676 (2017).
McArthur, A. G. et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 57, 3348â3357 (2013).
Suarez, N. M. et al. Superiority of transcriptional profiling over procalcitonin for distinguishing bacterial from viral lower respiratory tract infections in hospitalized adults. J. Infect. Dis. 212, 213â222 (2015).
Tsalik, E. L. et al. Host gene expression classifiers diagnose acute respiratory illness etiology. Sci. Transl. Med. 8, 322ra311 (2016).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. 5, eabd1554 (2020).
Zhou, Z. et al. Heightened innate immune responses in the respiratory tract of COVID-19 patients. Cell Host Microbe 27, 883â890 (2020).
Ong, E. Z. et al. A dynamic immune response shapes COVID-19 progression. Cell Host Microbe 27, 879â882 (2020).
Zhang, L. et al. Characterization of antibiotic resistance and host-microbiome interactions in the human upper respiratory tract during influenza infection. Microbiome 8, 39 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357â359 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15â21 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511â515 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139â140 (2010).
Anders, S., Pyl, P. T. & Huber, W. HTSeqâa Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166â169 (2015).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44â57 (2009).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078â2079 (2009).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674â1676 (2015).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Wang, Y. et al. GSA: genome sequence archive. Genomics Proteom. Bioinform. 15, 14â18 (2017).
Members, C.-N. & Partners Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2021. Nucleic Acids Res. 49, D18âD28 (2021).
Acknowledgements
The authors would like to thank Vazyme Biotech in Nanjing, China, for assistance in library procedure optimization and providing library preparation kits. In addition, the authors would like to thank National Center for Protein Sciences at Peking University in Beijing, China, for assistance with experiments. Part of the analysis was performed on the High Performance Computing Platform of the Center for Life Science (Peking University). This work was supported by International Innovation Resource Cooperation Project, Beijing Municipal Science and Technology Commission (Z201100008320024 to C.Y.), the National Natural Science Foundation of China (31861143026, 91940304 and 21825701 to C.Y.) and Epidemic Prevention and Control Special Project, Peking University.
Author information
Authors and Affiliations
Contributions
C.Y. and D.L. conceived the project; C.Y., D.L., H.W., and D.W. supervised the project; B.L., Y.Y., and L.D. designed the experiments together and wrote the manuscript; B.L., Y.Y., and L.H. performed experiments with the help of C.W. and R.W.; B.L. and Y.Y. performed the bioinformatics analysis; J.Y. collected the clinical samples; Y.L., J.C., D.Y., M.Z., and X.D. participated in discussion.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have filed a patent application related to TRACE-seq (patent application number: 202011520074.X).
Additional information
Publisherâs note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the articleâs Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the articleâs Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, B., Yan, Y., Dong, L. et al. Integrated characterization of SARS-CoV-2 genome, microbiome, antibiotic resistance and host response from single throat swabs. Cell Discov 7, 19 (2021). https://doi.org/10.1038/s41421-021-00248-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41421-021-00248-3
This article is cited by
-
Potential transmission chains of variant B.1.1.7 and co-mutations of SARS-CoV-2
Cell Discovery (2021)