Abstract
Pathogenic autoreactive antibodies that may be associated with life-threatening coronavirus disease 2019 (COVID-19) remain to be identified. Here, we show that self-assembled genome-scale libraries of full-length proteins covalently coupled to unique DNA barcodes for analysis by sequencing can be used for the unbiased identification of autoreactive antibodies in plasma samples. By screening 11,076 DNA-barcoded proteins expressed from a sequence-verified human ORFeome library, the method, which we named MIPSA (for Molecular Indexing of Proteins by Self-Assembly), allowed us to detect circulating neutralizing type-I and type-III interferon (IFN) autoantibodies in five plasma samples from 55 patients with life-threatening COVID-19. In addition to identifying neutralizing type-I IFN-α and IFN-Ï autoantibodies and other previously known autoreactive antibodies in patient plasma, MIPSA enabled the detection of as yet unidentified neutralizing type-III anti-IFN-λ3 autoantibodies that were not seen in healthy plasma samples or in convalescent plasma from ten non-hospitalized individuals with COVID-19. The low cost and simple workflow of MIPSA will facilitate unbiased high-throughput analyses of proteinâantibody, proteinâprotein and proteinâsmall-molecule interactions.
Similar content being viewed by others
Main
An unbiased analysis of antibody-binding specificities can provide insights into states of health and disease. We and others have used programmable phage-display libraries to identify novel autoantibodies, to characterize antiviral immunity and to profile allergen-specific IgE antibodies1,2,3,4,5. Although phage display has been useful for these and many other applications, most proteinâprotein, proteinâantibody and proteinâsmall-molecule interactions require a degree of conformational structure that is not captured by bacteriophage-displayed peptide libraries. Profiling conformational protein interactions at proteome scale has traditionally relied on protein microarray technologies. Protein microarrays, however, tend to suffer from high per-assay cost, and from a myriad of technical artefacts, including those associated with the high-throughput expression and purification of proteins, the spotting of proteins onto a solid support, the drying and rehydration of arrayed proteins and the readout of slides imaged via scanning fluorescence imaging6,7. Alternative approaches to protein-microarray production and storage have been developed (such as nucleic acid-programmable protein array, NAPPA8, or single-molecule polymerase chain reaction (PCR)-linked in vitro expression, SIMPLEX9). However, a robust, scalable and cost-effective alternative is lacking.
To overcome the limitations associated with the array-based profiling of full-length proteins, we previously established a methodology, which we named ParalleL Analysis of Translated Open reading frames (PLATO), that uses ribosome display of open reading frame (ORF) libraries10. Ribosome display relies on the in vitro translation of messenger RNAs that lack stop codons, stalling ribosomes at the ends of mRNA molecules in a complex with the nascent proteins that they encode. PLATO suffers from several key limitations that have hindered its adoption. An ideal alternative is the covalent conjugation of proteins to short amplifiable DNA barcodes. Indeed, individually prepared DNA-barcoded antibodies and proteins have been employed successfully in a variety of applications11. One particularly attractive proteinâDNA-conjugation method involves the HaloTag system, which adapts a bacterial enzyme that forms an irreversible covalent bond with halogen-terminated alkane moieties12. Individual DNA-barcoded HaloTag fusion proteins have been shown to greatly enhance the sensitivity and dynamic range of autoantibody detection, compared with traditional enzyme-linked immunosorbent assay13. Scaling individual protein barcoding to entire ORFeome libraries would be immensely valuable yet formidable, owing to high costs and low throughput. A self-assembly approach could provide a much more efficient path to library production.
In this Article, we describe a molecular-display technology, which we named Molecular Indexing of Proteins by Self-Assembly (MIPSA), that overcomes key disadvantages of PLATO and other full-length protein-array technologies. MIPSA produces libraries of soluble full-length proteins, each uniquely identifiable via covalent conjugation to an amplifiable DNA barcode. Barcodes are introduced upstream of the ribosome-binding site (RBS). Partial reverse transcription (RT) of the in vitro transcribed RNA (IVT-RNA) creates a complementary DNA barcode, which is linked to a haloalkane-labelled RT primer. An N-terminal HaloTag fusion protein is encoded downstream of the RBS, such that in vitro translation results in the intra-complex (âcisâ) covalent coupling of the cDNA barcode to the HaloTag and its downstream ORF-encoded protein product. The resulting library of uniquely indexed full-length proteins can be used for inexpensive proteome-wide interaction studies, such as unbiased autoantibody profiling.
Coronavirus disease 2019 (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection, ranges from an asymptomatic course to life-threatening pneumonia and death. A causal link between autoimmunity and severe COVID-19 has been supported by multiple studies14,15. Although a diverse array of autoantibodies have been documented16, neutralizing type-I interferon (IFN) autoantibodies seem to play a particularly prominent role17,18. Here, we investigate the utility of MIPSA by searching for novel autoantibodies in the plasma of patients with severe COVID-19.
Results
Development of the MIPSA system
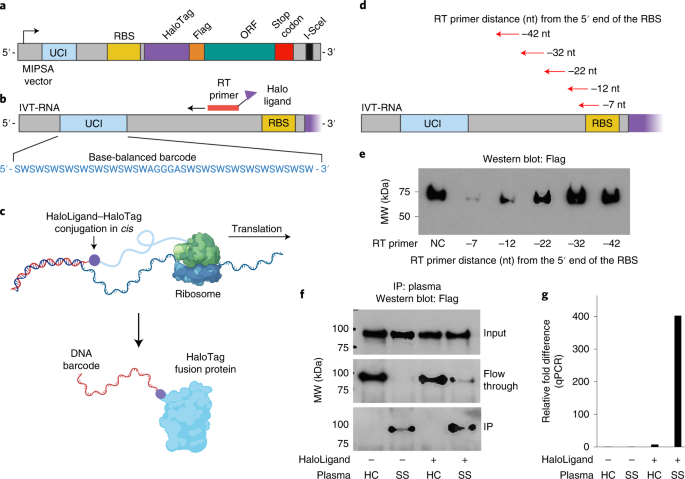
The MIPSA Gateway Destination vector for Escherichia coli cell-free translation contains the following key elements: a T7 RNA polymerase transcriptional start site, an isothermal unique clonal identifier (UCI) barcode sequence, an E. coli RBS, an N-terminal HaloTag fusion protein (891 nt), recombination sequences for ORF insertion and a homing endonuclease (I-SceI) site for plasmid linearization. A recombined ORF-containing pDESTâMIPSA plasmid is shown in Fig. 1a.
a, Schematic of the recombined pDESTâMIPSA vector with key components highlighted: UCI (blue), RBS (yellow), N-terminal HaloTag (purple), FLAG epitope (orange), ORF (green) and the I-SceI restriction endonuclease site (black) for vector linearization. b, Schematic showing IVT-RNA from the vector template shown in a. Isothermal base-balanced UCI sequence: (SW)18âAGGGAâ(SW)18. c, Cell-free translation of the RNAâcDNA shown in b. HaloTag protein forms a covalent bond with the HaloLigand-conjugated UCI-containing cDNA in cis during translation. d, RT primer positions tested for impact on translation. e, α-FLAG western blot analysis of translation in presence of RT primers depicted in d (NC, negative control, no RT primer). f, Western blot analysis of TRIM21 protein translated from RNA carrying the UCI-cDNA primed from the â32 position, either conjugated (+) or not (â) with the HaloLigand, IPed with healthy control (HC) or Sjögrenâs syndrome (SS) plasma. g, qPCR analysis of the immunoprecipitated TRIM21 UCI. Fold difference is by comparison with the HaloLigand (â) HC IP.
We first sought to establish a library of pDESTâMIPSA plasmids containing stochastic, isothermal UCIs located between the transcriptional start site and the RBS. A degenerate oligonucleotide pool was synthesized, comprising melting temperature (Tm) balanced sequences: (SW)18âAGGGAâ(SW)18, where S represents an equal mix of C and G, while W represents an equal mix of A and T (Fig. 1b). We reasoned that this inexpensive pool of sequences would (1) provide sufficient complexity (236 ~ 7âÃâ1010) for unique ORF labelling, (2) amplify without distortion and (3) serve as ORF-specific forward and reverse quantitative PCR (qPCR) primer binding sites for measurement of individual UCIs of interest. The degenerate oligonucleotide pool was amplified by PCR, restriction cloned into the MIPSA destination vector and transformed into E. coli (Methods). About 800,000 transformants were scraped off selection plates to obtain the pDESTâMIPSA UCI plasmid library. ORFs encoding the housekeeping protein glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and a known autoantigen, tripartite motif containing-21 (TRIM21, commonly known as Ro52) were separately recombined into the pDESTâMIPSA UCI plasmid library. Individually barcoded GAPDH and TRIM21 clones were isolated, sequenced and used in the following experiments.
The MIPSA procedure involves RT of the UCI using a succinimidyl ester (O2)-haloalkane (HaloLigand)-conjugated RT primer (Supplementary Fig. 1). The bound RT primer should not interfere with the assembly of the E. coli ribosome and initiation of translation but should be sufficiently proximal such that coupling of the HaloLigandâHaloTagâprotein complex might hinder additional rounds of translation (Fig. 1b,c). We tested a series of RT primers that anneal at distances ranging from â42 nucleotides to â7 nucleotides relative to the 5â² end of the RBS (Fig. 1d). On the basis of the yield of protein product from mRNA saturated with primers at these differing locations, we selected the â32 position as it did not interfere with translation efficiency (Fig. 1e). In contrast, RT from primers located within 20 nucleotides of the RBS diminished or abolished protein translation, in agreement with the estimated footprint of assembled 70S E. coli ribosomes, which have been shown to protect an average of 24 nucleotides of mRNA, with a range of 15 to 40 nucleotides.19
We next assessed the ability of SuperScript IV to perform RT from a primer labelled with the HaloLigand at its 5â² end, and the ability of the HaloTagâTRIM21 protein to form a covalent bond with the HaloLigand-conjugated primer during the translation reaction. HaloLigand conjugation and purification followed previously established methods. (Methods and Supplementary Fig. 1)20. Either an unconjugated RT primer or a HaloLigand-conjugated RT primer was used for RT of the barcoded HaloTagâTRIM21 mRNA. The translation product was then immunocaptured (immunoprecipitated) with plasma from a healthy donor or plasma from a TRIM21 autoantibody-positive patient with Sjögrenâs syndrome (SS), using protein-A- and protein-G-coated magnetic beads. The SS plasma efficiently immunoprecipitated the TRIM21 protein, regardless of RT primer conjugation, but only pulled down the TRIM21 UCI when the HaloLigand-conjugated primer was used in the RT reaction (Fig. 1f,g).
Assessing levels of cis versus trans UCI barcoding
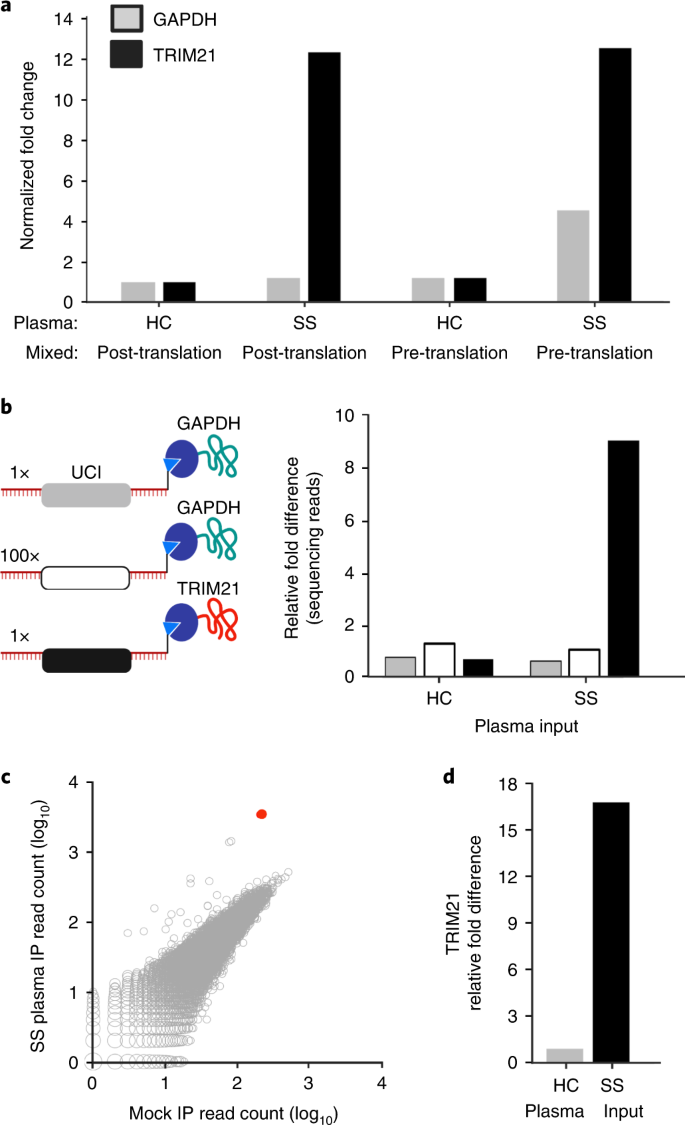
While the previous experiment indicated that, indeed, the HaloLigand does not impede RT priming and that the HaloTag can form a covalent bond with the HaloLigand during the translation reaction, it did not elucidate the amount of cis (intra-complex, desirable) versus trans (inter-complex, undesirable) HaloTagâUCI conjugation (Supplementary Fig. 2). Here, âintra-complexâ is defined as conjugation to the UCI that is associated with the same RNA molecule encoding the protein. To measure the amount of cis and trans HaloTagâUCI conjugation, GAPDH and TRIM21 mRNAs were separately reverse transcribed (using HaloLigand-conjugated primer) and then either mixed 1:1 or kept separate for in vitro translation. As expected, translation of the mixture produced roughly equivalent amounts of each protein compared with the individual translations (Supplementary Fig. 3). SS plasma specifically immunoprecipitated TRIM21 protein regardless of translation condition (Supplementary Fig. 3, immunoprecipitated fraction). However, we noted that while the SS IPs contained high levels of the TRIM21 UCI, as intended, more of the GAPDH UCI was pulled down by the SS plasma compared with that by the healthy control (HC) plasma when the mRNA was mixed before translation. This indicates that indeed some amount of trans barcoding occurs (Fig. 2a). We estimate that ~50% of the protein is cis-barcoded, with the remaining 50% trans-barcoded protein equally conjugated to both UCIs. Thus, in this two-component system, 25% of the TRIM21 protein is conjugated to the GAPDH UCI (Supplementary Fig. 2).
a, IVT-RNA encoding TRIM21 or GAPDH with their distinct UCI barcodes was translated before or after mixing at a 1:1 ratio. qPCR analysis of the IPs using UCI-specific primers, reported as fold change versus IP with healthy control (HC) plasma, which is set to 1, when the IVT-RNA was mixed post-translation. Sjögrenâs syndrome, SS. b, IVT-RNA encoding TRIM21 (black UCI) and GAPDH (grey UCI) were mixed 1:1 into a background of 100-fold excess GAPDH (white UCI) and then translated as a mock library. Sequencing analysis of the IPs, reported as fold change versus the HC IP of the 100à GAPDH. c, hORFeome MIPSA library containing spiked-in TRIM21, immunoprecipitated with SS plasma and compared with average of eight mock IPs (no plasma input). The TRIM21 UCI is shown in red. d, Relative fold difference of TRIM21 UCI in SS versus HC IPs, determined by sequencing.
In the setting of a complex library, even if ~50% of each protein is trans barcoded, this side product should be associated with a low level of randomly sampled UCIs. We tested this using a mock MIPSA library, composed of 100-fold excess of a second GAPDH clone, which was combined with a 1:1 mixture of the first GAPDH and TRIM21 clones (Fig. 2b).
Establishing and deconvoluting a stochastically barcoded human ORFeome MIPSA library
The sequence-verified human ORFeome (hORFeome) v8.1 is composed of 12,680 clonal ORFs mapping to 11,437 genes in the Gateway Entry plasmid (pDONR223)21. Five subpools of the library were created, each composed of ~2,500 similarly sized ORFs. Each of the five subpools was separately recombined into the pDESTâMIPSA UCI plasmid library and transformed to obtain approximately tenfold ORF coverage (~25,000 clones per subpool). Each subpool was assessed via Bioanalyzer electrophoresis, sequencing of ~20 colonies and Illumina sequencing of the combined superpool. The TRIM21 plasmid was spiked into the superpooled hORFeome library at 1:10,000âcomparable to a typical library member. The SS immunoprecipitation (IP) experiment was then performed on the hORFeome MIPSA library, using sequencing as a readout. The read counts from all UCIs in the library, including the spiked-in TRIM21, are shown for the SS IP versus the average of eight mock IPs in Fig. 2c. Reassuringly, the SS autoantibody-dependent enrichment of TRIM21 (17-fold) was similar to the model system (Fig. 2d). See âInformatic analysis of MIPSA sequencing dataâ in Methods for a description of the analytical pipeline for sequencing data.
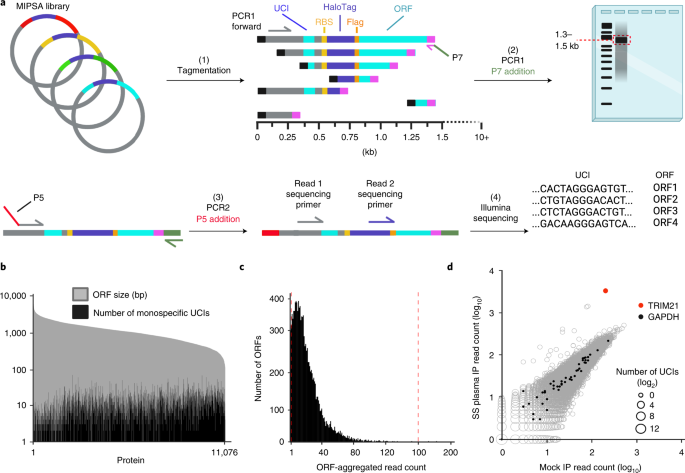
Next, we established a system for creating a UCIâORF look-up dictionary, using tagmentation and sequencing (Fig. 3a). Sequencing the 5â² 50 nt of the ORF inserts detected 11,076 of the 11,887 unique 5â² 50 nt library sequences. Of the 153,161 UCIs detected, 82.9% (126,975) were found to be associated with a single ORF (termed âmonospecific UCIâ). Each ORF was uniquely associated with a median of 9 (ranging from 0 to 123) monospecific UCIs (Fig. 3b). Importantly, an ensemble of monospecific UCIs with consistent behaviour can provide additional, strong support for the reactivity of their associated ORF. We noted a weak, inverse correlation between UCI number and ORF size, which most likely reflects the less efficient recombination of larger ORF-containing plasmids in the pooled recombination reactions. After aggregation of the read counts corresponding to each ORF, over 99% of the represented ORFs were present within a tenfold difference of the median ORF abundance (Fig. 3c). Taken together, these data indicate that we established a uniform library of 11,076 stochastically indexed human ORFs and defined a look-up dictionary for downstream analyses. Figure 3d shows UCI read counts of an SS IP versus the average of eight mock IPs and the 47 dictionary-decoded GAPDH monospecific UCIs (corresponding to two GAPDH isoforms present in the hORFeome library) appearing along the yâ=âx diagonal as expected. To avoid ambiguity, any UCI associated with more than a single ORF was excluded from further analyses.
a, (1) Tagmentation randomly inserts adapters into the MIPSA vector library. (2) Using a PCR1 forward primer and the reverse primer of the tagmentation-inserted adapter, DNA fragments are amplified and size selected to be ~1.5âkb, which captures the 5â² terminus of the ORF. (3) These fragments are amplified with a P5-containing PCR2 forward primer and a P7 reverse primer. (4) Illumina sequencing is used to read the UCI and the ORF from the same fragment, thus enabling their association in the dictionary. b, The number of monospecific UCIs is shown for each member of the pDESTâMIPSA hORFeome library, superimposed on the length of the ORFs. c, Histogram of ORF representations in the library according to their aggregated UCI-associated read counts. Vertical red lines show ±10à the median UCI-associated read count. d, IP of hORFeome MIPSA library using Sjögrenâs syndrome (SS) plasma is compared with the average of eight mock IPs. Sequencing read count for each UCI is plotted. UCIs associated with the two GAPDH isoforms (filled black) and spiked-in TRIM21 (red) are indicated.
Unbiased MIPSA analysis of autoantibodies associated with severe COVID-19
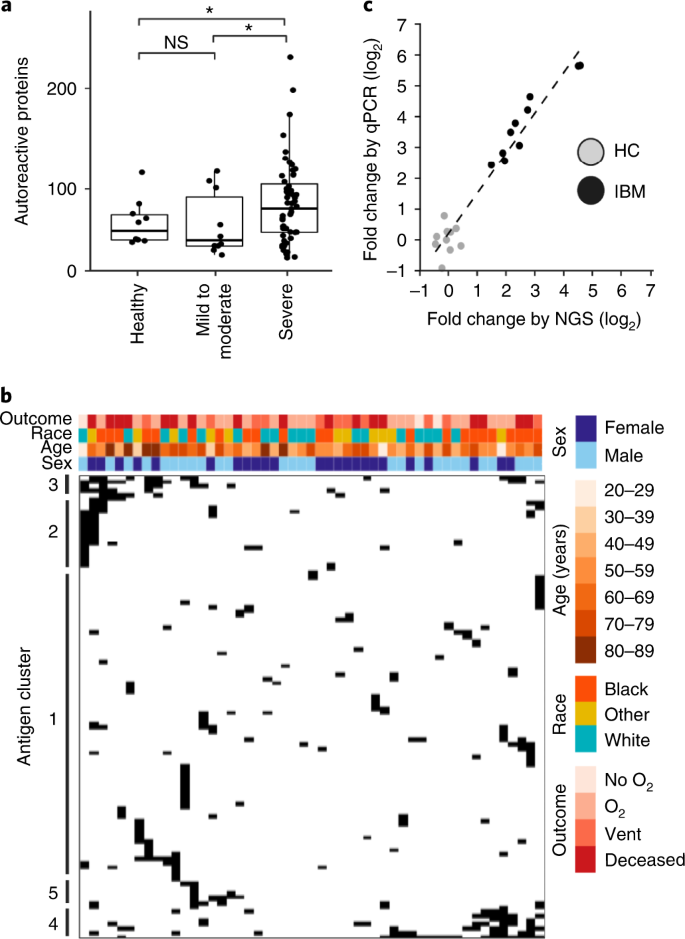
Several recent reports have described elevated autoantibody reactivities in patients with severe COVID-19 (refs. 22,23,24,25,26). We therefore used MIPSA with the human ORFeome library for unbiased identification of autoreactivities in the plasma of 55 patients with severe COVID-19, defined here only on the basis of hospital admission, since the availability of clinical meta-data was incomplete. For comparison, we used MIPSA to detect autoreactivities in plasma from ten healthy donors and ten COVID-19 convalescent plasma donors who had not been hospitalized (Supplementary Table 1). As we have done previously for phage immunoprecipitation sequencing (PhIP-seq) analyses, each sample was compared with a set of eight âmock IPsâ, which contained all reaction components except for plasma, and was run on the same plate. Comparison with mock IPs accounts for bias in the library and background binding. The informatic pipeline used to detect antibody-dependent reactivity (Methods) yielded a median of five (ranging from two to nine) false-positive UCI hits per mock IP. IPs using plasma from patients with severe COVID-19, however, yielded a mean of 83 reactive proteins among patients with severe COVID-19, which was significantly more than the mean of 64 reactive proteins among healthy pre-pandemic controls and significantly more than the mean of 62 reactive proteins among recovered individuals after mild to moderate COVID-19 (Pâ=â0.02 and Pâ=â0.05, respectively, one tailed t-test; Fig. 4a).
a, Box plots showing total numbers of autoreactive proteins in plasma from healthy controls (median 69.0 ± standard deviation (s.d.) 36.3), patients with mild-to-moderate COVID-19 (60.5 ± 44.7) or patients with severe COVID-19 (106.0 ± 67.6). Boxes indicate quartiles, and asterisks indicate Pâ=â0.02 and Pâ=â0.05 from a one-tailed t-test to compare means. NS, not significant. b, Hierarchal cluster map of all proteins represented by at least two reactive UCIs in at least one severe COVID-19 plasma, but no more than one control (healthy or mild-to-moderate COVID-19 plasma). c, MIPSA analysis of autoantibodies in ten patients with inclusion body myositis (IBM) and ten healthy controls (HC), using the hORFeome library. Fold change of immunoprecipitated NT5C1A, measured both as UCI-qPCR fold change (relative to average of ten HCs) and as sequencing fold change (relative to mock IPs).
We next examined proteins in the severe COVID-19 IPs that had at least two reactive UCIs (in the same IP) that were reactive in at least one severe patient and that were not reactive in more than one control (healthy or mild-to-moderate convalescent plasma). Proteins were excluded if they were reactive in a single patient with severe COVID-19 and a single control. The 103 proteins that met these criteria are shown in the cluster map of Fig. 4b. Fifty-one of the 55 patients with severe COVID-19 exhibited reactivity to at least one of these proteins. We noted co-occurring protein reactivities in multiple individuals, the vast majority of which lack homology by protein sequence alignment. Supplementary Table 2 provides summary statistics about these reactive proteins, including whether they are previously defined autoantigens according to the human autoantigen database AAgAtlas 1.0 (ref. 27). Supplementary dataset provides the patient versus UCI fold change data used to construct the cluster map.
One notable autoreactivity cluster (Supplementary Table 2, cluster 5) includes 5â²-nucleotidase, cytosolic 1A (NT5C1A), which is highly expressed in skeletal muscle and is the most well-characterized autoantibody target in inclusion body myositis (IBM). Multiple UCIs linked to NT5C1A were significantly increased in 3 of the 55 patients with severe COVID-19 (5.5%). NT5C1A autoantibodies have been reported in up to 70% of patients with IBM1, in ~20% of patients with SS and in up to ~5% of healthy donors.28 The prevalence of NT5C1A reactivity in the severe COVID-19 cohort is therefore not necessarily elevated. However, we wondered whether MIPSA would be able to reliably distinguish between healthy donor and IBM plasma on the basis of NT5C1A reactivity. We tested plasma from ten healthy donors and ten patients with IBM, the latter of whom were selected on the basis of NT5C1A seropositivity determined by PhIP-seq1. The clear separation of patients from controls in this independent cohort suggests that MIPSA may indeed have utility in clinical diagnostic testing using either UCI-specific qPCR or library sequencing, which were tightly correlated readouts (Fig. 4c).
Type-I and type-III IFN-neutralizing autoantibodies in patients with severe COVID-19
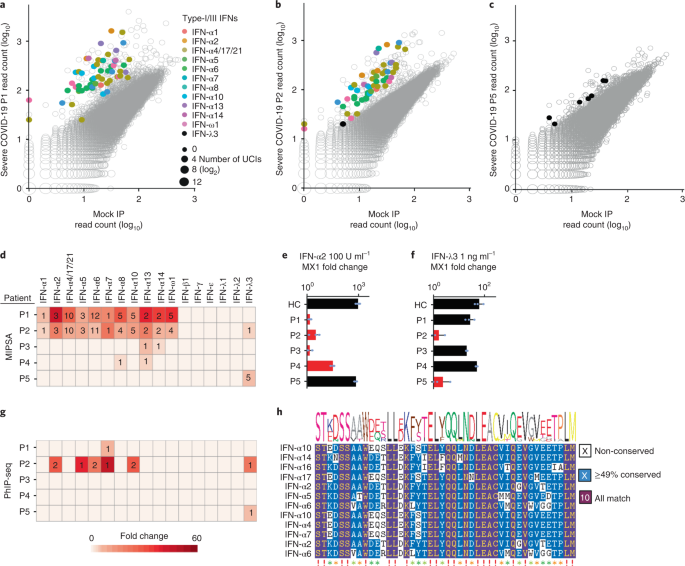
Neutralizing autoantibodies targeting type-I interferons alpha (IFN-α) and omega (IFN-Ï) have been associated with severe COVID-19 (refs. 16,23,29). All type-I IFNs except IFN-α16 are represented in the human MIPSA ORFeome library and annotated in the look-up dictionary. IFN-α4, IFN-α17 and IFN-α21 are indistinguishable by the first 50 nucleotides of their encoding ORF sequences and thus analysed as a single ORF. Two of the patients with severe COVID-19 (P1 and P2) in this cohort (3.6%) exhibited dramatic type-I IFN autoreactivity (49 and 46 type-I IFNs UCIs, across 11 distinct IFN-α and IFN-Ï ORFs; Fig. 5a,b). The extensive co-reactivity of these proteins is probably attributable to their sequence homology (Supplementary Fig. 4). By requiring at least two reactive IFN UCIs to be considered positive, we identified two additional severe COVID-19 plasma samples (P3 and P4) with detectable levels of IFN-α reactivity, each with only two reactive IFN-α UCIs. Fifty percent of these four patients with autoreactive IFN-α died, versus about 30% of the remaining cohort. Interestingly, one additional plasma sample (P5) precipitated no UCIs from any type-I or type-II IFNs, but five UCIs from the type-III IFN, IFN-λ3 (Fig. 5c,d). This patient also died of COVID-19. No additional IFN autoreactivities were detected among the patients with severe COVID-19. None of the healthy or non-hospitalized COVID-19 controls was positive for two or more IFN UCIs.
aâc, Scatterplots highlighting reactive IFN UCIs for three patients with severe COVID-19. d, Summary of IFN reactivity detected in 5 of 55 individuals with severe COVID-19. Hit fold-change values (colour of cell) and the number of reactive UCIs (number in cell) are provided. e,f, Recombinant IFN-α2 or IFN-λ3 neutralizing activity of the same patients shown in d. Plasma were pre-incubated with 100 U mlâ1 of IFN-α2 or 1âng mlâ1 of IFN-λ3 before incubation with A549 cells. Fold changes of the IFN stimulated gene, MX1, were calculated by RTâqPCR relative to unstimulated cells (nâ=â3 for each of the five patient samples). For e, samples labelled HC, P1, P2, P3, P4 and P5 have mean and s.d. values of 977.7â±â253.7, 1.39â±â0.2, 3.1â±â1.8, 1.4â±â0.5, 32â±â7.2 and 741.1â±â121, respectively. For f, samples labelled HC, P1, P2, P3, P4 and P5 have mean and s.d. values of 64.5â±â26.8, 28.2â±â12.1, 1.6â±â1.0, 20.3â±â1.4, 52.4â±â5.1 and 2.3â±â2.0, respectively. GAPDH was used as a housekeeping control gene for normalization. Red bars indicate which samples were found to contain the corresponding anti-IFN antibodies using MIPSA. g, PhIP-seq analysis of IFN autoantibodies in the five patients of d (row and column orders maintained). Hit fold-change values (colour of cell) and the number of reactive peptides (number in cell) are provided. h, Epitopefindr analysis of the PhIP-seq reactive type-I IFN 90-aa peptides.
We further assessed the performance of MIPSA using P2 plasma, which neutralizes both type-I and type-III IFNs. MIPSA was run on P2 plasma in triplicate, yielding a high level of assay reproducibility (Supplementary Fig. 5a,b), both in consistent detection of hits and low coefficients of variation (mean 22%). We next determined the linearity of the assay by diluting P2 plasma tenfold into healthy plasma and then performing MISPA again. The results demonstrate a consistent decrease in signal among the reactivities (mean of 5.4-fold for reactive IFNs) and loss of detection of some hits, particularly of ORFs with single reactive UCIs.
Incubation of A549 human adenocarcinomatous lung epithelial cells with 100 U mlâ1 IFN-α or 1âng mlâ1 of IFN-λ3 for 4âh in serum-free medium results in a robust upregulation of the IFN-response gene MX1 by ~1,000-fold and ~100-fold, respectively. Pre-incubation of IFN-α2 with plasma P1, P2 or P3 completely abolished MX1 upregulation (Fig. 5e). The plasma with the weakest IFN-α reactivity by MIPSA, P4, only partially neutralized the cytokine. Neither HC nor P5 plasma had any effect on the response of A549 cells to IFN-α2 treatment. However, pre-incubation of the IFN-λ3 cytokine with the MIPSA-positive plasma, P2 and P5, ablated the IFN response (Fig. 5f). None of the other plasma (HC, P1, P3 or P4) had any effect on the response of A549 cells to IFN-λ3. By comparison against titration curves using IFN-α2 and IFN-λ3 monoclonal antibodies, a serial titration using patient P2 plasma in triplicate indicated circulating levels of these autoantibodies to be ~20âμg mlâ1 and ~100âng mlâ1, respectively (Supplementary Fig. 6). MIPSA analysis of the serially diluted IFN-α2 mAb revealed broad IFN-α cross-recognition but mutually exclusive binding of the monoclonal antibodies to the appropriate type-I or type-III IFN (Supplementary Fig. 7a,b). Importantly, we noted that loss of MIPSA detection sensitivity corresponded to the same or greater plasma dilutions at which IFN-α2 and IFN-λ3 neutralization activities were also lost. Finally, the titre of P2âs autoantibodies exhibited at least a ten-fold preference for IFN-λ3 neutralization over IFN-λ1 neutralization (Supplementary Fig. 8). In summary, MIPSA-based autoantibody profiling of this severe COVID-19 cohort identified strongly neutralizing IFN-α autoantibodies in 7.3% of patients and strongly neutralizing IFN-λ3 autoantibodies in 3.6% of patients, with a single patient (1.8%) harbouring both autoreactivities.
We then determined whether PhIP-seq with a 90-amino-acid (aa) human peptidome library30 might also detect IFN autoantibodies in this cohort. PhIP-seq detected IFN-α reactivity in plasma from P1 and P2, although to a much lesser extent (Fig. 5g). The two weaker IFN-α reactivities detected by MIPSA in the plasma of P3 and P4 were both missed by PhIP-seq. PhIP-seq identified a single additional weakly IFN-α reactive sample, which was negative by MIPSA (not shown). Both technologies detected type-III IFN autoreactivity (directed exclusively at IFN-λ3). PhIP-seq data were used to narrow the location of a dominant epitope in these type-I and type-III IFN autoantigens (Fig. 5h for IFN-α; amino acid position 45â135 for IFN-λ3).
We next wondered about the prevalence of the previously unreported IFN-λ3 autoreactivity in the general population and whether it might be increased among patients with severe COVID-19. PhIP-seq was previously used to profile the plasma of 423 HCs, none of whom was found to have detectable IFN-λ3 autoreactivity.31 These data suggest that IFN-λ3 autoreactivity is likely to be more frequent among individuals with severe COVID-19. Therefore, neutralizing IFN-λ3 autoantibodies may be involved in a pathogenic mechanism contributing to life-threatening COVID-19 in a subset of patients.
Discussion
We have described a molecular-display technology for full-length proteins that provides key advantages over protein microarrays and alternative techniques (such as PLATO). MIPSA uses self-assembly to produce a library of proteins, linked to relatively short (158 nt) single-stranded cDNA barcodes via the 25-kDa HaloTag domain. This compact barcoding approach will probably have many applications not accessible to alternative display formats with bulky linkage cargoes (such as yeast, bacteria, viruses, phages, ribosomes, mRNAs and cDNAs). Indeed, individually conjugating minimal DNA barcodes to proteins, especially antibodies and antigens, have already proven useful in several settings, including CITE-seq (Cellular Indexing of Transcriptomes and Epitopes by Sequencing)32, LIBRA-seq (linking B cell receptor to antigen specificity through sequencing)33 and related methodologies34. At proteome scale, MIPSA will enable unbiased analyses of proteinâantibody, proteinâprotein and proteinâsmall-molecule interactions, as well as studies of post-translational modifications, such as hapten-modification studies35 or protease-activity profiling36. Key advantages of MIPSA include its high throughput, low cost, simple sequencing-library preparation, inherent compatibility with PhIP-seq and the stability of the proteinâDNA complexes (important for the manipulation and storage of display libraries). Importantly, MIPSA can be adopted by standard molecular biology laboratories, since it does not require specialized training or instrumentation (but does require access to a high-throughput DNA-sequencing instrument or facility).
Autoantibodies detected in patients with severe COVID-19 using MIPSA
Neutralizing IFN-α/Ï autoantibodies have been described in patients with severe COVID-19 disease and are presumed to be pathogenic.23 These likely pre-existing autoantibodies, which occur very rarely in the general population, block restriction of viral replication in cell culture and are thus likely to interfere with disease resolution. This discovery paved the way to identifying a subset of individuals at risk for life-threatening COVID-19 and proposed therapeutic use of IFN-β in this population of patients. In our study, MIPSA identified two individuals with extensive reactivity to the entire family of IFN-α cytokines. Indeed, plasma from both individuals, plus two individuals with weaker IFN-α reactivity detected by MIPSA, robustly neutralized recombinant IFN-α2 in a lung adenocarcinomatous cell culture model.
Type-III IFNs (IFN-λ, also known as IL-28/29) are cytokines with potent anti-viral activities that act primarily at barrier sites. The IFN-λR1/IL-10RB heterodimeric receptor for IFN-λ is expressed on lung epithelial cells and is important for the innate response to viral infection. Previous studies in mice determined that IFN-λ diminished pathogenicity and suppressed replication of influenza viruses, respiratory syncytial virus, human metapneumovirus and severe acute respiratory syndrome coronavirus (SARS-CoV-1)37. It has been proposed that IFN-λ exerts much of its antiviral activity in vivo via stimulatory interactions with immune cells, rather than through induction of the antiviral cell state38. However, IFN-λ has been found to robustly restrict SARS-CoV-2 replication in primary human bronchial epithelial cells39, primary human airway epithelial cultures40 and primary human intestinal epithelial cells41. Collectively, these studies suggest multifaceted mechanisms by which neutralizing IFN-λ autoantibodies may exacerbate SARS-CoV-2 infections.
Among 55 patients with severe COVID-19, MIPSA detected two individuals with IFN-λ3 reactive autoantibodies. The same autoreactivities were also detected using PhIP-seq. We tested the IFN-λ3 neutralizing capacity of these patientsâ plasma, observing near-complete ablation of the cellular response to the recombinant cytokine (Fig. 5f). These data suggest that IFN-λ3 autoreactivity is a potentially pathogenic mechanism contributing to severe COVID-19 disease.
In one study, type-III IFN neutralizing antibodies were not detected among a cohort of 101 individuals with type-I IFN autoantibodies tested.23 In our study, one of the four IFN-α autoreactive individuals (P2, a 22-year-old male) also harboured autoantibodies that neutralized IFN-λ3. It is possible that this co-reactivity is extremely rare and thus not represented in the aforementioned 101-patient study. Alternatively, it is possible that the differing assay conditions exhibit different detection sensitivity. Whereas in the previous study cultured A549 cells were incubated with IFN-λ3 at 50âng mlâ1 without plasma pre-incubation, we cultured A549 cells with IFN-λ3 at 1âng mlâ1 after pre-incubation with plasma for 1 h. Their readout of STAT3 phosphorylation may also provide different detection sensitivity compared with the upregulation of MX1 expression. A larger study is needed to determine the true frequency of these reactivities in patients with severe COVID-19 and matched controls. Here, we report detection of strongly neutralizing IFN-α and IFN-λ3 autoantibodies in 4 (7.3%) and 2 (3.6%) individuals, respectively, in a cohort of 55 patients with severe COVID-19. IFN-λ3 autoantibodies were not detected via PhIP-seq in a larger cohort of 423 HCs collected before the pandemic.
Exogenously administered type-III IFNs have been proposed as a therapeutic for SARS-CoV-2 infection40,42,43,44,45,46, and there are currently three ongoing clinical trials to test PEGylated IFN-λ1 for efficacy in reducing morbidity and mortality associated with COVID-19 (ClinicalTrials.gov Identifiers NCT04343976, NCT04534673 and NCT04344600). One recently completed double-blind, placebo-controlled trial, NCT04354259, reported a significant reduction by 2.42âlog copies per millilitre of SARS-CoV-2 at day 7 among patients with mild-to-moderate COVID-19 in the outpatient setting (Pâ=â0.0041)47. Future studies will determine whether anti-IFN-λ3 autoantibodies are pre-existing or arise in response to SARS-CoV-2 infection and how often they also cross-neutralize IFN-λ1. On the basis of neutralization data from P2 (Supplementary Fig. 8) and sequence alignment of IFN-λ1 and IFN-λ3 (~29% homology, Supplementary Fig. 4), cross-neutralization is expected to be rare, raising the possibility that patients with neutralizing IFN-λ3 autoantibodies may derive benefit from PEGylated IFN-λ1 treatment.
While clusters of uncharacterized autoreactivities were observed in multiple individuals, it is not clear what role, if any, they may play in severe COVID-19. In larger-scale studies, we expect that patterns of co-occurring reactivity, or reactivities towards proteins with related biological functions, may ultimately define new autoimmune syndromes associated with severe COVID-19.
Complementarity of MIPSA and PhIP-seq
Display technologies frequently complement one another but may not be amenable to routine simultaneous use. MIPSA is more likely than PhIP-seq to detect antibodies directed at conformational epitopes on proteins expressed well in vitro. This was exemplified by the robust detection of IFN-α autoantibodies via MIPSA, which were less sensitively detected via PhIP-seq. PhIP-seq, on the other hand, is more likely to detect antibodies directed at less conformational epitopes contained within proteins that are either absent from an ORFeome library or cannot be expressed well in cell-free lysate. Because MIPSA and PhIP-seq naturally complement one another in these ways, we designed the MIPSA UCI amplification primers to be the same as those we have used for PhIP-seq. As the UCIâprotein complex is stableâeven in phage preparationsâMIPSA and PhIP-seq can readily be performed together in a single reaction, using a single set of amplification and sequencing primers. The compatibility of these two display modalities lowers the barrier to leveraging their synergy.
Variations of the MIPSA system
A key aspect of MIPSA involves the conjugation of a protein to its associated UCI in cis, compared with another library memberâs UCI in trans. Here, we have used covalent conjugation via the HaloTag/HaloLigand system, but others could work as well. For instance, the SNAP-tag (a 20âkDa mutant of the DNA repair protein O6-alkylguanine-DNA alkyltransferase) forms a covalent bond with benzylguanine (BG) derivatives48. BG could thus be used to label the RT primer in place of the HaloLigand. A mutant derivative of the SNAP-tag, the CLIP-tag, binds O2-benzylcytosine derivatives, which could also be adapted to MIPSA49.
The rate of HaloTag maturation and ligand binding is critical to the relative yield of cis versus trans UCI conjugation. A previous study determined that the rate of HaloTag protein production is about fourfold higher than the rate of HaloTag functional maturation50. Considering a typical protein size is <1,000 amino acids in the ORFeome library, these data predict that most proteins should be released from the ribosome before HaloTag maturation, and thus before cis HaloLigand conjugation could occur, thereby favoring unwanted trans barcoding. However, we observed ~50% of proteinâUCI conjugates are formed in cis, thereby enabling excellent assay performance in the setting of a complex library. During optimization experiments, we found the rate of cis barcoding to be slightly improved by excluding release factors from the translation mix, which stalls ribosomes on their stop codons and allows HaloTag maturation to continue in proximity to its UCI. Alternative approaches to promote controlled ribosomal stalling could include stop codon removal/suppression or use of a dominant negative release factor. Ribosome release could then be induced via addition of the chain terminator puromycin.
Since UCI cDNAs are formed on the 5â² UTR of the IVT-RNA, eukaryotic ribosomes would be unable to scan from the 5â² cap to the initiating Kozak sequence. The MIPSA system described here is therefore incompatible with cap-dependent eukaryotic cell-free translation systems. If cap-dependent translation is desired, however, two alternative methods could be developed. First, the current 5â² UCI system could be used if an internal ribosome entry site were to be placed between the RT primer and the Kozak sequence. Second, the UCI could instead be introduced at the 3â² end of the RNA, provided that the RT was prevented from extending into the ORF. In an extension of eukaryotic MIPSA, RNAâcDNA hybrids could potentially be transfected into living cells or tissues, where UCI-protein formation could take place in situ, enabling many additional applications.
The ORF-associated UCIs can be embodied in a variety of ways. Here, we have stochastically assigned indexes to the human ORFeome at ~10Ã representation. This approach has two main benefits: first, a single degenerate oligonucleotide pool is low cost; second, multiple independent measurements are reported by the ensemble of UCIs associated with each ORF. We have designed our library of UCIs with uniform GC content and, thus, uniform PCR amplification efficiency. For simplicity, we have opted not to incorporate unique molecular identifiers into the RT primer, but this approach is compatible with MIPSA UCIs and may potentially enhance quantitation. One disadvantage of stochastic indexing is the potential for ORF dropout and, thus, the need for relatively high UCI representation; this increases the depth of sequencing required to quantify each UCI and, thus, the overall per-sample cost. A second disadvantage is the requirement to construct a UCIâORFeome matching dictionary. With short-read sequencing, we were unable to disambiguate a fraction of the library, composed mostly of alternative isoforms. Using a long-read sequencing technology, such as PacBio or Oxford Nanopore Technologies, instead of or in addition to short-read sequencing technology could surmount incomplete disambiguation. As opposed to stochastic barcoding, individual UCIâORF cloning is possible but costly and cumbersome. However, a smaller UCI set would provide the advantage of lower per-assay sequencing cost. We have previously developed a methodology to clone ORFeomes using Long Adapter Single Stranded Oligonucleotide (LASSO) probes51. LASSO cloning of ORFeome libraries thus naturally synergizes with MIPSA-based applications.
MIPSA readout via qPCR
A useful feature of appropriately designed UCIs is that they can also serve as qPCR readout probes. The degenerate UCIs that we have designed and used here (Fig. 1b) comprise 18 nt base-balanced forward and reverse primer binding sites. The low cost and rapid turnaround time of a qPCR assay can thus be leveraged in combination with MIPSA. For example, incorporating assay quality-control measures, such as TRIM21 IP, can be used to qualify a set of samples before a costlier sequencing run. Troubleshooting and optimization can similarly be expedited by employing qPCR as a readout, rather than relying exclusively on next-generation DNA sequencing (NGS). qPCR testing of specific UCIs may theoretically also provide enhanced sensitivity compared with sequencing and may be more amenable to analysis in a clinical setting.
Outlook
MIPSA is a protein-display technology that has key advantages over alternative approaches. It has properties that complement techniques such as PhIP-seq, and the MIPSA ORFeome libraries can be conveniently screened in the same reactions with phage-display libraries. The MIPSA protocol requires cap-independent cell-free translation, but future adaptations may overcome this limitation. Applications for MIPSA-based studies include proteinâprotein, proteinâantibody and proteinâsmall-molecule interaction studies, as well as analyses of post-translational modifications. We used MIPSA to detect known autoantibodies and to discover neutralizing IFN-λ3 autoantibodies, among many other potentially pathogenic autoreactivities (Supplementary Table 2) that may contribute to life-threatening COVID-19 in a subset of at-risk individuals.
Methods
MIPSA destination vector construction
The MIPSA vector was constructed using the Gateway pDEST15 vector as a backbone. A gBlock fragment (Integrated DNA Technologies) encoding the RBS, Kozak sequence, N-terminal HaloTag fusion protein and FLAG tag, followed by an attR1 sequence, was cloned into the parent plasmid. A 150âbp poly(A) sequence was also added after attR2 site. The TRIM21 and GAPDH ORF sequences used for characterizing and optimizing the two-component system included native stop codons that were retained in the final MIPSA construct.
UCI barcode library construction
A 41 nt barcode oligo was generated within a gBlock Gene Fragment (Integrated DNA Technologies) with alternating mixed bases (S: G/C; W: A/T) to produce the following sequence: (SW)18âAGGGAâ(SW)18. The sequences flanking the degenerate barcode incorporated the standard PhIP-seq PCR1 and PCR2 primer binding sites52. Eighteen nanograms of the starting UCI library was used to run 40 cycles of PCR to amplify the library and incorporate BglII and PspxI restriction sites. The MIPSA vector and amplified UCI library were then digested with the restriction enzymes overnight, column purified and ligated at 1:5 vector-to-insert ratio. The ligated MIPSA vector was used to transform electrocompetent One Shot ccdB 2 T1R cells (Thermo Fisher Scientific). Six transformation reactions yielded ~800,000 colonies to produce the pDESTâMIPSA UCI library.
Human ORFeome recombination into the pDESTâMIPSA UCI plasmid library
A total of 150âng of each pENTRâhORFeome subpool (L1âL5) from hORFeome v8.1 was individually combined with 150âng of the pDESTâMIPSA UCI library plasmid and 2âμl of Gateway LR Clonase II mix (Life Technologies) for a total reaction volume of 10âμl. The reaction was incubated overnight at 25â°C. The entire reaction was transformed into 50âμl of One Shot OmniMAX 2 T1R chemical competent E. coli (Life Technologies). In aggregate, the transformations yielded ~120,000 colonies, which is approximately tenfold the complexity of the hORFeome v8.1. Colonies were collected and pooled by scraping, followed by purification of the barcoded pDESTâMIPSA hORFeome plasmid DNA (human ORFeome MIPSA library) using the Qiagen Plasmid Midi Kit (Qiagen). The human hORFeome v8.1 collection was cloned without stop codons; the displayed proteins may therefore contain poly-lysine C-termini resulting from translation of the polyA tail. A more recent version of the MIPSA destination vector includes a stop codon in frame with recombined ORFs.
HaloLigand conjugation to RT oligo and HPLC purification
One-hundred micrograms of a 5â² amine modified oligo HL-32_ad (Supplementary Table 3) was incubated with 75âμl (17.85âμg μlâ1) of the HaloTag succinimidyl ester (O2) (Promega Corporation), the HaloLigand, in 0.1âM sodium borate buffer for 6âh at room temperature following ref. 20. Then, 3âM NaCl and ice-cold ethanol was added at 10% (v/v) and 250% (v/v), respectively, to the labelling reaction and incubated overnight at â80â°C. The reaction was centrifuged for 30âmin at 12,000g. The pellet was rinsed once in ice-cold 70% ethanol and air dried for 10âmin.
HaloLigand-conjugated RT primer was HPLC purified using a Brownlee Aquapore RP-300 7âU, 100 à 4.6âmm column (Perkin Elmer) using a two-buffer gradient of 0â70% CH3CN/MeCN (100âmM triethylamine acetate to acetonitrile) over 70âmin. Fractions corresponding to labelled oligo were collected and lyophilized (Supplementary Fig. 1). Oligos were resuspended at 1âμM (15.4âng µlâ1) and stored at â80â°C.
MIPSA library IVT-RNA preparation
The human ORFeome MIPSA library plasmid (4âμg) was linearized with I-SceI restriction endonuclease (New England Biolabs) overnight. The product was column purified with the NucleoSpin Gel and PCR Clean Up Kit (Macherey-Nagel). A 40âμl in vitro transcription reaction using the HiScribe T7 High Yield RNA Synthesis Kit (New England Biolabs) was used to transcribe 1âμg of the purified, linearized pDESTâMIPSA plasmid library. The product was diluted with 60âμl molecular biology grade water, and 1âμl of DNAse I was added. The reaction was incubated for another 15âmin at 37â°C. Then 50âμl of 1âM LiCl was added to the solution and incubated at â80â°C overnight. A centrifuge was cooled to 4â°C, and the RNA was spun at maximum speed for 30âmin. The supernatant was removed, and the RNA pellet washed with 70% ethanol. The sample was spun down at 4â°C for another 10âmin, and the 70% ethanol removed. The pellet was dried at room temperature for 15âmin and subsequently resuspended in 100âμl water. To preserve the sample, 1âμl of 40 U μlâ1 RNAseOUT Recombinant Ribonuclease Inhibitor (Life Technologies) was added.
MIPSA library IVT-RNA RT and translation
An RT reaction was prepared using SuperScript IV First-Strand Synthesis System (Life Technologies). First, 1âμl of 10âmM dNTPs, 1âμl of RNAseOUT (40 U μlâ1), 4.17âμl of the RNA library (1.5âμM) and 7.83âμl of the HaloLigand-conjugated RT primer (1âμM, Supplementary Table 3) were combined in a single 14âμl reaction and incubated at 65â°C for 5âmin followed by a 2-min incubation on ice. Four microlitres of 5à RT buffer, 1âμl of 0.1âM dithiothreitol and 1âμl of SuperScript IV RT Enzyme (200 U μlâ1) were added to the 14âμl reaction on ice and incubated for 20âmin at 42â°C. A single 20âμl RT reaction received 36âμl of RNAClean XP beads (Beckman Coulter) and was incubated at room temperature for 10âmin. The beads were collected by magnet and washed five times with 70% ethanol. The beads were air dried for 10âmin at room temperature and resuspended in 7âμl of 5âmM TrisâHCl, pH 8.5. The product was analysed with spectrophotometry to measure the RNA yield. A translation reaction was set up on ice using the PURExpress ÎRibosome Kit (New England Biolabs)53. The reaction was modified such that the final concentration of ribosomes was 0.3âμM. For each 10âμl translation reaction, 4.57âμl of the RT reaction was added to 4âμl Solution A, 1.2âμl Factor Mix and 0.23âμl ribosomes (13.3âμM). This reaction was incubated at 37â°C for 2 h, diluted to a total volume of 45âμl with 35âμl 1à PBS and used immediately or stored at â80â°C after addition of glycerol to a final concentration of 25% (v/v).
IP of the translated MIPSA hORFeome library
Five microlitres of plasma, diluted 1:100 in PBS, is mixed with the 45âμl of diluted MIPSA library translation reaction (see above) and incubated overnight at 4â°C with gentle agitation. For each IP, a mixture of 5âμl of Protein A Dynabeads and 5âμl of Protein G Dynabeads (Life Technologies) was washed three times in two times their original volume with 1à PBS. The beads were then resuspended in 1à PBS at their original volume and added to each IP. The antibody capture proceeded for 4âh at 4â°C. Beads were collected on a magnet and washed three times in 1à PBS, changing tubes or plates between washes. The beads were then collected and resuspended in a 20âμl PCR master mix containing the T7-Pep2_PCR1_F forward and the T7-Pep2_PCR1_Râ+âad_min reverse primers (Supplementary Table 3) and Herculase-II (Agilent). PCR cycling was as follows: an initial denaturing and enzyme activation step at 95â°C for 2âmin, followed by 20 cycles of 95â°C for 20âs, 58â°C for 30âs and 72â°C for 30âs. The final extension step was performed at 72â°C for 3âmin. Two microlitres of the PCR1 amplification product was used as input to a 20âμl dual-indexing PCR reaction with the PhIP_PCR2_F forward and the PhIP_PCR2_R reverse primers, each containing 10 nt barcodes (i5 and i7, respectively). PCR cycling was as follows: an initial denaturing step at 95â°C for 2âmin, followed by 20 cycles of 95â°C for 20âs, 58â°C for 30âs and 72â°C for 30âs. The final extension step was performed at 72â°C for 3âmin. i5/i7 indexed libraries were pooled and column purified (NucleoSpin columns, Takara). Libraries were sequenced on an Illumina NextSeq 500 using a 1 à 50 nt SE or 1 à 75 nt SE protocol. MIPSA_i5_NextSeq_SP and Standard_i7_SP primers were used for i5/i7 sequencing (Supplementary Table 3). The output was demultiplexed using i5 and i7 without allowing any mismatches.
For quantification of MIPSA experiments by qPCR, the PCR1 product (above) was analysed as follows. A total of 4.6âμl of 1:1,000 dilution of the PCR1 reaction was added to 5âμl of Brilliant III Ultra Fast 2à SYBR Green Mix (Agilent), 0.2âμl of 2âμM reference dye and 0.2âμl of 10âμM forward and reverse primer mix (specific to the target UCI). PCR cycling was as follows: an initial denaturing step at 95â°C for 2âmin, followed by 45 cycles of 95â°C for 20âs and 60â°C for 30 s. Following completion of thermocycling, amplified products were subjected to melt-curve analysis. The qPCR primers for MIPSA IP experiments were BT2_F and BT2_R for TRIM21, BG4_F and BG4_R for GAPDH and NT5C1A_F and NT5C1A_R for NT5C1A (Supplementary Table 3).
Plasma samples
All samples were collected from subjects who met protocol eligibility criteria, as described below. All studies protected the rights and privacy of the study participants and were approved by their respective institutional review boards for original sample collection and subsequent analyses.
Pre-pandemic and HC plasma samples
All human samples were collected before 2017 at the National Institutes of Health (NIH) Clinical Center under the Vaccine Research Centerâs (VRC)/National Institutes of Allergy and Infectious Diseases (NIAID)/NIH protocol âVRC 000: Screening Subjects for HIV Vaccine Research Studiesâ (NCT00031304) in compliance with NIAID institutional review board-approved procedures.
COVID-19 convalescent plasma from non-hospitalized patients
Eligible non-hospitalized COVID-19 convalescent plasma donors were contacted by study personnel, as previously described54. All donors were at least 18 years old and had a confirmed diagnosis of SARS-CoV-2 by detection of RNA in a nasopharyngeal swab sample. Basic demographic information (age, sex, race and hospitalization with COVID-19) was obtained from each donor; initial diagnosis of SARS-CoV-2 and date of diagnosis were confirmed by medical chart review.
Severe COVID-19 plasma samples
The study cohort was defined as inpatients who had (1) a confirmed RNA diagnosis of COVID-19 from a nasopharyngeal swab sample, (2) survival to death or discharge and (3) remnant specimens in the Johns Hopkins COVID-19 Remnant Specimen Biorepository, an opportunity sample that includes 59% of Johns Hopkins Hospital patients with COVID-19 and 66% of patients with length of stay â¥3 days55,56. Patient outcomes were defined by the World Health Organization COVID-19 disease severity scale. Samples from patients with severe COVID-19 that were included in this study were obtained from 17 patients who died, 13 who recovered after being ventilated, 22 who required oxygen to recover and 3 who recovered without supplementary oxygen. This study was approved by the Johns Hopkins University institutional review board (IRB00248332 and IRB00273516), with a waiver of consent because all specimens and clinical data were de-identified by the Core for Clinical Research Data Acquisition of the Johns Hopkins Institute for Clinical and Translational Research; the study team had no access to identifiable patient data.
SS and IBM plasma samples
SS samples were collected under protocol NA_00013201. All patients were >18 years old and gave informed consent. Samples from patients with IBM were collected under protocol IRB00235256. All patients met ENMC 2011 diagnostic criteria57 and provided informed consent.
Immunoblot analysis
Laemmli buffer containing 5% β-mercaptoethanol was added to samples, boiled for 5âmin and analysed on NuPage 4â12% Bis-Tris polyacrylamide gels (Life Technologies). Following transfer to PVDF membranes, blots were blocked in 20âmM Tris-buffered saline, pH 7.6, containing 0.1% Tween 20 (TBST) and 5% (wt/vol) non-fat dry milk for 30âmin at room temperature. Blots were subsequently incubated overnight at 4â°C with primary anti-FLAG antibody (#F3165, MilliporeSigma) at 1:2,000 (v/v), followed by a 4-h incubation at room temperature in anti-mouse IgG, HRP-linked secondary antibody (#7076, Cell Signaling) at 1:4,000 (v/v).
Construction of the UCIâORF dictionary
The Nextera XT DNA Library Preparation kit (Illumina) was used for tagmentation of 150âng of the pDESTâMIPSA hORFeome plasmid library to yield the optimal size distribution centred around 1.5âkb. Tagmented libraries were amplified using Herculase-II (Agilent) with T7-Pep2_PCR1_F forward and Nextera Index 1 Read primer. PCR cycling was as follows: an initial denaturing step at 95â°C for 2âmin, followed by 30 cycles of 95â°C for 20âs, 53.5â°C for 30âs and 72â°C for 30âs. A final extension step was performed at 72â°C for 3âmin. PCR reactions were run on a 1% agarose gel followed by excision of ~1.5âkb products and purification using the NucleoSpin Gel and PCR Clean-up columns (Macherey-Nagel). The purified product was then amplified for another ten cycles with PhIP_PCR2_F forward and P7.2 reverse primers (for a list of primer sequences, see Supplementary Table 3). The product was gel purified and sequenced on a MiSeq (Illumina) using the T7-Pep2.2_SP_subA primer for read 1 and the MISEQ_MIPSA_R2 primer for read 2. Read 1 was 60âbp long to capture the UCIs. The first index read, I1, was substituted with a 50âbp read into the ORF. I2 was used to identify the i5 index for sample demultiplexing.
The hORFeome v8.1 DNA sequences were truncated to the first 50 nt, and the ORF names corresponding to non-unique sequences were concatenated with a â|â delimiter. The demultiplexed output of the 50 nt R2 (ORF) read from an Illumina MiSeq was aligned to the truncated human ORFeome v8.1 library using the Rbowtie2 package with the following parameters: optionsâ=âââa âvery-sensitive-localâ58. The unique FASTQ identifiers were then used to extract corresponding sequences from the 60âbp R1 (UCI) read. Those sequences were then trimmed using the 3â² anchor ACGATA, and sequences that did not have the anchor were removed. Additionally, any trimmed R1 sequences that had fewer than 18 nucleotides were removed. The ORF sequences that still had a corresponding UCI post-filtering were retained using the FASTQ identifier. The names of ORFs that had the same UCI were concatenated with a â&â delimiter, and this final dictionary was used to generate a FASTA alignment file composed of ORF names and UCI sequences.
Informatic analysis of MIPSA sequencing data
Illumina output FASTQ files were trimmed using the constant ACGAT anchor sequence following all UCI sequences. Next, perfect match alignment was used to map the trimmed sequences to their linked ORFs via the UCIâORF look-up dictionary. A read count matrix was constructed, in which rows correspond to individual UCIs and columns correspond to samples. We next used the edgeR software package59, which, using a negative binomial model, compares the signal detected in each sample against a set of negative control (âmockâ) IPs that were performed without plasma, to return a fold-change estimate and a test statistic for each UCI in every sample, thus creating fold-change and âlog10P matrices. By comparison of EdgeR output data from replicate IPs, we established that significantly enriched UCIs (âhitsâ) should require a read count of at least 15, a P value less than 0.001 and a fold change of at least 3. Hit fold-change matrices report the fold-change value for âhitsâ and report a â1â for UCIs that are not hits.
Protein sequence similarity
To evaluate sequence homology among proteins in the hORFeome v8.1 library, a blastp alignment was used to compare each protein sequence against all other library members (parameters â-outfmt 6 -evalue 100 -max_hsps 1 -soft_masking false -word_size 7 -max_target_seqs 100000â). To evaluate sequence homology among reactive peptides in the human 90-aa phage display library, the epitopefindr (brandonsie.github.io/epitopefindr) software was employed.
PhIP-seq analyses
PhIP-seq was performed according to a previously published protocol.52 Briefly, 0.2âμl of each plasma was individually mixed with the 90-aa human phage library and immunoprecipitated using protein-A- and protein-G-coated magnetic beads. A set of six to eight mock IPs (no plasma input) were run on each 96-well plate. Magnetic beads were resuspended in PCR master mix and subjected to thermocycling. A second PCR reaction was employed for sample barcoding. Amplicons were pooled and sequenced on an Illumina NextSeq 500 instrument using a 1âÃâ50ânt SE or 1âÃâ75ânt SE protocol. PhIP-seq with the human library was used to characterize autoantibodies in a collection of plasma from HCs. For fair comparison with the severe COVID-19 cohort, we first determined the minimum sequencing depth that would have been required to detect the IFN-λ3 reactivity in both of the positive individuals. We then considered only the 423 datasets from the healthy cohort with sequencing depth greater than this minimum threshold. None of these 423 individuals was found to be reactive to any peptide from IFN-λ3.
Type-I/III IFN neutralization assay
IFN-α2 (catalogue no. 11100-1), IFN-λ1 (catalogue no. 1598-IL-025) and IFN-λ3 (catalogue no. 5259-IL-025) were purchased from R&D Systems. Twenty microlitres of plasma were incubated for 1âh at room temperature with either 100 U mlâ1 IFN-α2 or 1âng mlâ1 IFN-λ3, and 180âμl DMEM in a total volume of 200âµl before addition into 7.5âÃâ104 A549 cells in 48-well tissue culture plates. After 4-h incubation, the cells were washed with 1à PBS and cellular mRNA was extracted and purified using RNeasy Plus Mini Kit (Qiagen). Six hundred nanograms of extracted mRNA was reverse transcribed using the SuperScript III First-Strand Synthesis System (Life Technologies) and diluted tenfold for qPCR analysis on a QuantStudio 6 Flex System (Applied Biosystems). PCR consisted of 95â°C for 3âmin, followed by 45 cycles of the following: 95â°C for 15âs and 60â°C for 30âs. MX1 expression was chosen as a measure of cell stimulation by the IFNs, and the relative mRNA expression was normalized to GAPDH expression. The qPCR primers for GAPDH and MX1 were obtained from Integrated DNA Technologies (Supplementary Table 3). Anti-hIFN-α2-IgG (catalogue no. mabg-hifna-3) and anti-hIL-28b-IgG (catalogue no. mabg-hil28b-3) were purchased from InvivoGen. Manufacturerâs note about mabg-hifna-3: âThis antibody reacts with hIFN-α1, hIFN-α2, hIFN-α5, hIFN-α8, hIFN-α14, hIFN-α16, hIFN-α17 and hIFN-α21; it reacts very weakly with hIFN-α4 and IFN-α10; it does not react with hIFN-α6 or hIFN-α7.â The manufacturerâs note about mabg-hil28b-3: âReacts with human IL-28A and human IL-28B.â
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data supporting the results in this study are available within the paper and its Supplementary Information. The raw and analysed datasets are available from the corresponding author on request. Source data are provided with this paper.
Code availability
The MIPSAlign package for alignment and UCIâORF matching, implemented in R v 4.0.2, is available on GitHub at https://github.com/jgunn123/MIPSAlign.
References
Larman, H. B. et al. Cytosolic 5â²-nucleotidase 1A autoimmunity in sporadic inclusion body myositis. Ann. Neurol. 73, 408â418 (2013).
Xu, G. J. et al. Viral immunology. Comprehensive serological profiling of human populations using a synthetic human virome. Science 348, aaa0698 (2015).
Shrock, E. et al. Viral epitope profiling of COVID-19 patients reveals cross-reactivity and correlates of severity. Science https://doi.org/10.1126/science.abd4250 (2020).
Monaco, D. R. et al. Profiling serum antibodies with a pan allergen phage library identifies key wheat allergy epitopes. Nat. Commun. 12, 379 (2021).
Venkataraman, T. et al. Analysis of antibody binding specificities in twin and SNP-genotyped cohorts reveals that antiviral antibody epitope selection is a heritable trait. Immunity 55, 174â184.e5 (2022).
Kingsmore, S. F. Multiplexed protein measurement: technologies and applications of protein and antibody arrays. Nat. Rev. Drug Discov. 5, 310â320 (2006).
Kodadek, T. Protein microarrays: prospects and problems. Chem. Biol. 8, 105â115 (2001).
Ramachandran, N., Hainsworth, E., Demirkan, G. & LaBaer, J. On-chip protein synthesis for making microarrays. Methods Mol. Biol. 328, 1â14 (2006).
Rungpragayphan, S., Yamane, T. & Nakano, H. SIMPLEX: single-molecule PCR-linked in vitro expression: a novel method for high-throughput construction and screening of protein libraries. Methods Mol. Biol. 375, 79â94 (2007).
Zhu, J. et al. Protein interaction discovery using parallel analysis of translated ORFs (PLATO). Nat. Biotechnol. 31, 331â334 (2013).
Liszczak, G. & Muir, T. W. Nucleic acid-barcoding technologies: Converting DNA sequencing into a broad-spectrum molecular counter. Angew. Chem. Int. Ed. Engl. 58, 4144â4162 (2019).
Los, G. V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373â382 (2008).
Yazaki, J. et al. HaloTag-based conjugation of proteins to barcoding-oligonucleotides. Nucleic Acids Res. 48, e8 (2020).
Liu, Y., Sawalha, A. H. & Lu, Q. COVID-19 and autoimmune diseases. Curr. Opin. Rheumatol. 33, 155â162 (2021).
Knight, J. S. et al. The intersection of COVID-19 and autoimmunity. J. Clin. Invest. https://doi.org/10.1172/JCI154886 (2021).
Wang, E. Y. et al. Diverse functional autoantibodies in patients with COVID-19. Nature 595, 283â288 (2021).
Bastard, P. et al. Autoantibodies neutralizing type I IFNs are present in ~4% of uninfected individuals over 70 years old and account for ~20% of COVID-19 deaths. Sci Immunol. https://doi.org/10.1126/sciimmunol.abl4340 (2021).
Abers, M. S. et al. Neutralizing type-I interferon autoantibodies are associated with delayed viral clearance and intensive care unit admission in patients with COVID-19. Immunol. Cell Biol. 99, 917â921 (2021).
Mohammad, F., Green, R. & Buskirk, A. R. A systematically-revised ribosome profiling method for bacteria reveals pauses at single-codon resolution. eLife https://doi.org/10.7554/eLife.42591 (2019).
Gu, L. et al. Multiplex single-molecule interaction profiling of DNA-barcoded proteins. Nature 515, 554â557 (2014).
Yang, X. et al. A public genome-scale lentiviral expression library of human ORFs. Nat. Methods 8, 659â661 (2011).
Consiglio, C. R. et al. The immunology of multisystem inflammatory syndrome in children with COVID-19. Cell 183, 968â981 e967 (2020).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science https://doi.org/10.1126/science.abd4585 (2020).
Zuo, Y. et al. Prothrombotic autoantibodies in serum from patients hospitalized with COVID-19. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.abd3876 (2020).
Casciola-Rosen, L. et al. IgM anti-ACE2 autoantibodies in severe COVID-19 activate complement and perturb vascular endothelial function. JCI Insight. 7, 1â18 (2022).
Woodruff, M. C., Ramonell, R. P., Lee, F. E. & Sanz, I. Relaxed peripheral tolerance drives de novo autoreactivity in severe COVID-19. medRxiv https://doi.org/10.1101/2020.10.21.20216192 (2021).
Wang, D. et al. AAgAtlas 1.0: a human autoantigen database. Nucleic Acids Res. 45, D769âD776 (2017).
Lloyd, T. E. et al. Cytosolic 5â²-nucleotidase 1A as a target of circulating autoantibodies in autoimmune diseases. Arthritis Care Res. 68, 66â71 (2016).
Gupta, S., Nakabo, S., Chu, J., Hasni, S. & Kaplan, M. J. Association between anti-interferon-alpha autoantibodies and COVID-19 in systemic lupus erythematosus. medRxiv https://doi.org/10.1101/2020.10.29.20222000 (2020).
Xu, G. J. et al. Systematic autoantigen analysis identifies a distinct subtype of scleroderma with coincident cancer. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1615990113 (2016).
Venkataraman, T. et al. Analysis of antibody binding specificities in twin and SNP-genotyped cohorts reveals that antiviral antibody epitope selection is a heritable trait. Immunity 55, 174â184 e175 (2022).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865â868 (2017).
Setliff, I. et al. High-throughput mapping of B cell receptor sequences to antigen specificity. Cell 179, 1636â1646 e1615 (2019).
Saka, S. K. et al. Immuno-SABER enables highly multiplexed and amplified protein imaging in tissues. Nat. Biotechnol. 37, 1080â1090 (2019).
Roman-Melendez, G. D. et al. Citrullination of a phage-displayed human peptidome library reveals the fine specificities of rheumatoid arthritis-associated autoantibodies. EBioMedicine 71, 103506 (2021).
Roman-Melendez, G. D., Venkataraman, T., Monaco, D. R. & Larman, H. B. Protease activity profiling via programmable phage display of comprehensive proteome-scale peptide libraries. Cell Syst. 11, 375â381 e374 (2020).
Mordstein, M. et al. Lambda interferon renders epithelial cells of the respiratory and gastrointestinal tracts resistant to viral infections. J. Virol. 84, 5670â5677 (2010).
Ank, N. et al. Lambda interferon (IFN-λ), a type III IFN, is induced by viruses and IFNs and displays potent antiviral activity against select virus infections in vivo. J. Virol. 80, 4501â4509 (2006).
Busnadiego, I. et al. Antiviral activity of type I, II, and III interferons counterbalances ACE2 inducibility and restricts SARS-CoV-2. mBio https://doi.org/10.1128/mBio.01928-20 (2020).
Vanderheiden, A. et al. Type I and type III interferons restrict SARS-CoV-2 infection of human airway epithelial cultures. J. Virol. https://doi.org/10.1128/JVI.00985-20 (2020).
Stanifer, M. L. et al. Critical role of type III interferon in controlling SARS-CoV-2 infection in human intestinal epithelial cells. Cell Rep. 32, 107863 (2020).
Galani, I. E. et al. Untuned antiviral immunity in COVID-19 revealed by temporal type I/III interferon patterns and flu comparison. Nat. Immunol. 22, 32â40 (2021).
Felgenhauer, U. et al. Inhibition of SARS-CoV-2 by type I and type III interferons. J. Biol. Chem. 295, 13958â13964 (2020).
OâBrien, T. R. et al. Weak induction of interferon expression by severe acute respiratory syndrome coronavirus 2 supports clinical trials of interferon-λ to treat early coronavirus disease 2019. Clin. Infect. Dis. 71, 1410â1412 (2020).
Andreakos, E. & Tsiodras, S. COVID-19: lambda interferon against viral load and hyperinflammation. EMBO Mol. Med. 12, e12465 (2020).
Prokunina-Olsson, L. et al. COVID-19 and emerging viral infections: The case for interferon lambda. J. Exp. Med. https://doi.org/10.1084/jem.20200653 (2020).
Feld, J. J. et al. Peginterferon lambda for the treatment of outpatients with COVID-19: a phase 2, placebo-controlled randomised trial. Lancet Respir. Med. https://doi.org/10.1016/S2213-2600(20)30566-X (2021).
Jongsma, M. A. & Litjens, R. H. Self-assembling protein arrays on DNA chips by auto-labeling fusion proteins with a single DNA address. Proteomics 6, 2650â2655 (2006).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128â136 (2008).
Samelson, A. J. et al. Kinetic and structural comparison of a proteinâs cotranslational folding and refolding pathways. Sci. Adv. 4, eaas9098 (2018).
Tosi, L. et al. Long-adapter single-strand oligonucleotide probes for the massively multiplexed cloning of kilobase genome regions. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-017-0092 (2017).
Mohan, D. et al. Publisher correction: PhIP-seq characterization of serum antibodies using oligonucleotide-encoded peptidomes. Nat. Protoc. 14, 2596 (2019).
Tuckey, C., Asahara, H., Zhou, Y. & Chong, S. Protein synthesis using a reconstituted cell-free system. Curr. Protoc. Mol. Biol. 108, 16 31 11â16 31 22 (2014).
Klein, S. L. et al. Sex, age, and hospitalization drive antibody responses in a COVID-19 convalescent plasma donor population. J. Clin. Invest. 130, 6141â6150 (2020).
Garibaldi, B. T. et al. Patient trajectories among persons hospitalized for COVID-19. Ann. Intern. Med. 174, 144 (2021).
Zyskind, I. et al. SARS-CoV-2 seroprevalence and symptom onset in culturally-linked Orthodox Jewish communities across multiple regions in the United States. JAMA Open Netw. 4, 1â9 (2021).
Rose, M. R. & Group, E. I. W. 188th ENMC International Workshop: inclusion body myositis, 2â4 December 2011, Naarden, The Netherlands. Neuromuscul. Disord. 23, 1044â1055 (2013).
Wei, Z., Zhang, W., Fang, H., Li, Y. & Wang, X. esATAC: an easy-to-use systematic pipeline for ATAC-seq data analysis. Bioinformatics 34, 2664â2665 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139â140 (2010).
Acknowledgements
This study was made possible by a Johns Hopkins University Provost Research Grant made through the Johns Hopkins COVID-19 Research Response Program, by a National Institute of General Medical Sciences (NIGMS) grant R01 GM127353 (to H.B.L. and B.P.), by a grant from the National Heart, Lung, and Blood Institute of the National Institutes of Health under award number K23HL151826 (to E.M.B.), by grants from the Sjögrenâs Syndrome Foundation and the Jerome L. Greene Foundation (to A.N.B. and H.B.L.) and by funding from the intramural research programs of the Vaccine Research Center, National Institute of Allergy and Infectious Diseases and the National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Institutes of Health. We thank S. Elledge for generously providing the human ORFeome library and the 90-aa human peptidome T7 phage display library. We thank R. Green, M. Catipovic, A. Buskirk, T. OâDonnell, P. Duggal and J. Markle for helpful discussions. We also thank J. Franklin and the Johns Hopkins Synthesis and Sequencing Core for HPLC purification of the HaloLigand conjugated RT-primer, as well as L. Orzolek and H. Hao of the Johns Hopkins Transcriptomics and Deep Sequencing Core Facility. We also thank C. Tuckey at New England Biolabs for assistance with Translation kits. The severe COVID-19 specimens used for this work were part of the Johns Hopkins Biospecimen Repository, which relies on the contribution of many patients, research teams and clinicians. We thank the members of the NIH Vaccine Research Center for pre-pandemic sample collection: B. Graham, L. Novick, J. Casazza, J. Ledgerwood, U. Sarwar, L. Chang, C. Starr Hendel, L. Holman, S. Plummer, P. Costner, I. Gorden, B. Larkin, F. Mendoza, J. Saudners, K. Zephir, M. E. Enama, G. Yamshchikov, I. Pittman and P. Williams. Parts of some figures were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
H.B.L. and J.J.C. conceived the project and oversaw all experiments. H.B.L., J.J.C., J.G. and P.S. wrote the manuscript. J.J.C., A.R., A.W. and H.-J.T. constructed the MIPSA destination vector. J.J.C., J.G. and P.S. performed the MIPSA experiments and IFN neutralization studies. J.G. and D.R.M. wrote the MIPSA computational pipelines. J.G., X.A.Z., D.R.M., W.R.M and Y.D. performed data analysis and edited the manuscript. B.P. and L.T. reviewed data and edited the paper. A.L.C. oversaw selection of remnant specimens from the Johns Hopkins COVID-19 Remnant Specimen Biorepository and edited the manuscript. E.M.B and A.A.R.T oversaw collection of COVID-19 convalescent plasma from non-hospitalized patients and edited the manuscript. A.N.B., M.R., I.Z., A.Z.R., J.I.S., S.J., T.L. and A.L.M were involved in the collection and analysis of data from clinical samples and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
H.B.L., J.J.C., J.G. and P.S. are listed as inventors on a patent application filed by Johns Hopkins University that covers the MIPSA technology. H.B.L. is a founder of Portal Bioscience, Alchemab and ImmuneID, and an advisor to TScan Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisherâs note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and tables.
Supplementary Dataset
MIPSA data matrix of hit fold changes, for UCIs of reactive proteins in patients with severe COVID-19.
Source data
Source Data Fig. 1
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Credle, J.J., Gunn, J., Sangkhapreecha, P. et al. Unbiased discovery of autoantibodies associated with severe COVID-19 via genome-scale self-assembled DNA-barcoded protein libraries. Nat. Biomed. Eng 6, 992â1003 (2022). https://doi.org/10.1038/s41551-022-00925-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-022-00925-y
This article is cited by
-
Mitochondrial dysfunction in long COVID: mechanisms, consequences, and potential therapeutic approaches
GeroScience (2024)
-
Human antibody profiling technologies for autoimmune disease
Immunologic Research (2023)
-
Autoantibodies Neutralizing Type I IFNs in the Bronchoalveolar Lavage of at Least 10% of Patients During Life-Threatening COVID-19 Pneumonia
Journal of Clinical Immunology (2023)