Advertisement
Research ArticleOncology Free access | 10.1172/JCI73351
MicroRNA-205 signaling regulates mammary stem cell fate and tumorigenesis
Chi-Hong Chao,1,2 Chao-Ching Chang,1,2 Meng-Ju Wu,1,2 How-Wen Ko,3,4,5 Da Wang,3 Mien-Chie Hung,3,4,6 Jer-Yen Yang,1,2 and Chun-Ju Chang1,2
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Chao, C. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Chang, C. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Wu, M. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Ko, H. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Wang, D. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Hung, M. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Yang, J. in: JCI | PubMed | Google Scholar
1Department of Basic Medical Sciences, College of Veterinary Medicine, and 2Center for Cancer Research, Purdue University, West Lafayette, Indiana, USA. 3Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA. 4The University of Texas Graduate School of Biomedical Sciences at Houston, Houston, Texas, USA. 5Department of Thoracic Medicine, Chang Gung Memorial Hospital, Chang Gung University College of Medicine, Taoyuan, Taiwan. 6The Center for Molecular Medicine and Graduate Institute of Cancer Biology, China Medical University, Taichung, Taiwan.
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Find articles by Chang, C. in: JCI | PubMed | Google Scholar
Authorship note: Chi-Hong Chao and Chao-Ching Chang contributed equally to this work.
Published June 9, 2014 - More info
J Clin Invest. 2014;124(7):3093–3106. https://doi.org/10.1172/JCI73351.
Copyright © 2014, American Society for Clinical Investigation
Received: September 26, 2013; Accepted: April 21, 2014
-
Abstract
Dysregulation of epigenetic controls is associated with tumorigenesis in response to microenvironmental stimuli; however, the regulatory pathways involved in epigenetic dysfunction are largely unclear. We have determined that a critical epigenetic regulator, microRNA-205 (miR-205), is repressed by the ligand jagged1, which is secreted from the tumor stroma to promote a cancer-associated stem cell phenotype. Knockdown of miR-205 in mammary epithelial cells promoted epithelial-mesenchymal transition (EMT), disrupted epithelial cell polarity, and enhanced symmetric division to expand the stem cell population. Furthermore, miR-205–deficient mice spontaneously developed mammary lesions, while activation of miR-205 markedly diminished breast cancer stemness. These data provide evidence that links tumor microenvironment and microRNA-dependent regulation to disruption of epithelial polarity and aberrant mammary stem cell division, which in turn leads to an expansion of stem cell population and tumorigenesis. This study elucidates an important role for miR-205 in the regulation of mammary stem cell fate, suggesting a potential therapeutic target for limiting breast cancer genesis.
-
Introduction
Cancer stem cells, a subpopulation of cancer cells that have acquired the stemness properties associated with normal stem cells, are considered to be the genesis of cancer and account for cancer initiation, progression, and recurrence (1). It has been shown that an enlarged cancer stem cell population is highly associated with tumor aggressiveness (2) and that, in response to microenvironmental stimuli, the cancer stem cell population can be expanded to drive cancer progression, potentially through dysregulation of genetic or epigenetic mechanisms (3). Therefore, it is important to understand the key regulatory mechanism of cancer stemness and to develop effective therapeutic strategies to eradicate the genesis of cancer.
NOTCH signaling components are frequently upregulated in invasive breast cancer (4). Upon interaction of the ligands (e.g., jagged1) with the NOTCH receptors, the intracellular domain of the NOTCH (NICD) is released from the cytoplasmic membrane to the nucleus through a cascade of proteolytic cleavage by the metalloprotease enzyme and γ-secretase, leading to transcriptional activation of the NOTCH target genes, such as HES1 (4). The NOTCH ligand jagged1 is known to be overexpressed in tumor cells as well as in the tumor stroma, and jagged1 expression within the stem cell niche plays a role in nurturing the hematopoietic, hepatic, and neural stem/progenitor cells (5, 6). Interestingly, a recent study also demonstrated that soluble jagged1 can be secreted from the tumor stroma to promote the cancer stem cell phenotype (7). However, the regulatory mechanism by which jagged1 signaling modulates cancer stem cell phenotypes remains to be elucidated.
micro-RNAs (miRNAs), small noncoding RNA molecules that suppress gene expression by interacting with the 3′ untranslated regions (3′ UTRs) of target messenger RNAs, regulate a myriad of biological processes, including the cell fate decision (8). A previous study has reported that microRNA-205 (miR-205) is one of the most significantly downregulated miRNAs in human breast tumors compared with normal tissues (9). Notably, low expression of miR-205 predicts a chemotherapy relapse in cancer patients who have triple-negative breast cancer (TNBC) (9), where a high content of the cancer stem cell population is enriched. It is intriguing that emerging in vitro studies reveal complex roles of miR-205 as either a tumor suppressor or an oncogene, depending on different cell contexts (10). Nonetheless, the role of miR-205 in breast cancer in vivo and the mechanism by which miR-205 is regulated during tumorigenesis still remain unclear.
This study reveals that jagged1, which was shown to be secreted by the tumor stroma (7), promotes the stemness phenotype through downregulating miR-205. A feedback regulatory loop of NOTCH/miR-205/ZEB1 signaling is uncovered as being critical for regulation of epithelial-mesenchymal transition (EMT) and polarity of stem cell division for maintaining the mammary epithelial homeostasis. Dysregulation of miR-205 expression leads to the mesenchymal phenotype, disrupted epithelial cell polarity, and expansion of the symmetrically self-renewing stem cell population, which further contribute to mammary tumorigenesis in vivo. Our findings elucidate a mechanism by which miR-205, serving as a master switch, coordinates the microenvironmental queue and its downstream signaling to control the tumor stem cell population, revealing important clinical implications for miR-205 in prediction and treatment of aggressive breast cancer by regulating tumor stemness.
-
Results
The ligand jagged1 suppresses miR-205 expression through HES1-mediated transcriptional repression. Accumulated evidence suggests that jagged1 signals from the stem/progenitor cell niche are critical for governing the cell fate decision (5, 6). A recent study also revealed that soluble jagged1, which is secreted from the tumor stroma, interacts with membrane-bound NOTCH receptors to activate NOTCH signaling and promote the cancer stem cell phenotype (7). To explore the potential epigenetic mechanism that could be involved in the regulation of this process, such as the miRNAs, we analyzed changes in the global miRNA expression profile in response to jagged1 treatment (active peptide mimicking the soluble jagged1 in ref. 7) using a genome-wide miRNA-PCR array consisting of 1,066 annotated miRNAs. miR-205 was identified as the most significantly downregulated miRNA (5.1-fold reduction, n = 3, P < 0.03) in the primary human mammary epithelial cells (HMEC) under jagged1 treatment (Figure 1A). Consistent with the result, the highest jagged1 expression was found persistently on the surface of human breast tumor cells that were in the proximity of the tumor-stroma junction region (Figure 1B), where miR-205 expression was significantly reduced (Figure 1B and Table 1; n = 68, P = 0.0001). To further gain insight into the regulatory mechanism of miR-205, we analyzed, using promoter analysis, the response elements of transcription factors located within the 3-kb region upstream of the transcription start site of MIR205. We found 2 putative HES1 response elements located within the MIR205 promoter that had high consensus scores (Figure 1C; matrix similarity score > 0.9, score = 1 as a perfect match). Since HES1 is a transcription repressor and a prime target downstream of activated jagged1-NOTCH signaling, it is reasonable to speculate that jagged1 treatment downregulates miR-205 expression through HES1-mediated transcription repression.
 Figure 1
Figure 1The ligand jagged1 significantly represses miR-205 to promote EMT and stem cell phenotype. (A) Expression levels of the most significantly altered miRNAs in HMECs treated with 40 μM jagged1 or the control peptide for 1 day using genome-wide miRNA-PCR array (n = 3, *P < 0.03). (B) Representative image showing a reverse correlation between expression of miR-205 and jagged1. Scale bars: 25 μm. S, stroma, T, tumor. (C) Diagram showing the promoter regions of MIR205 with the putative HES1 response elements (HES1-RE). (D) ChIP-PCR showing the percentage of the bound chromatin/input chromatin using HES1 antibody targeting HES1-binding elements (A and B) in BT549 breast cancer cells transfected with control or HES1 siRNA. IgG was used as a negative control (n = 3, *P < 0.05). (E) miR-205/mRNA expression levels of BT549 cells transfected with control or HES1 siRNA (n = 3, *P < 0.05). (F) Cell morphology of MCF12A cells stably expressing shRNA of the control vector (sh-Vec) and miR-205 (sh–miR-205). Scale bar: 20 μm. (G) miRNA and mRNA expression and (H) protein expression levels of sh-Vec and sh–miR-205–MCF12A cells. (I) The percentage of the CD24–CD44+ population (lower right quadrant) in sh-Vec and sh–miR-205 cells (n = 3, *P < 0.05). (J) The number of forming spheres per 1,000 cells generated from sh-Vec and sh–miR-205–MCF12A cells (n = 3, *P < 0.05). Scale bars: 100 μm, Error bars denote mean ± SD.
Indeed, real-time PCR data suggest that jagged1 treatment could induce HES1 gene expression to downregulate miR-205 (Supplemental Figure 1A; supplemental material available online with this article; doi:10.1172/JCI73351DS1). miR-205 has also been shown to regulate EMT through inhibiting ZEB1 (11), a transcription repressor of the epithelial cell marker E-cadherin. Consistently, our data showed that jagged1 activated ZEB1 and repressed E-cadherin expression (CDH1), which was reversed by forced expression of miR-205 (Supplemental Figure 1A). Moreover, knockdown (KD) of jagged1 gene expression using lentivirus-delivered shRNA (sh-JAG1) resulted in decreased HES1 expression, increased miR-205 expression, and a reduced level of ZEB1 (Supplemental Figure 1B).
To further validate the direct association of HES1 with the MIR205 promoter, we performed ChIP analysis on both HES1-binding elements within the MIR205 promoter region (elements A and B, Figure 1C) using an antibody specifically against HES1. The ChIP results revealed that HES1 was more significantly bound to element A (Figure 1D). KD of HES1 markedly diminished the amount of DNA that could be immunoprecipitated by the HES1 antibody (Figure 1D), and the diminished DNA was accompanied by increased miR-205 expression and a reduced level of ZEB1 (Figure 1E). Together, these data suggest that HES1, activated by jagged1-NOTCH signaling, inhibits miR-205 expression through direct binding to the specific promoter element of MIR205.
Downregulation of miR-205 expands mammary stem cell and tumor stem cell populations. Additionally, we found that jagged1 treatment significantly promoted the stem cell/tumor stem cell population stained for CD24–CD44+, an established stem cell/tumor stem cell surface marker, in primary HMECs, MCF12A mammary epithelial cells, and human primary breast tumor cells (PT) (Supplemental Figure 1C). We found that the percentage of the CD24–CD44+ population was inversely correlated with the endogenous levels of miR-205 in these cell lines, and the enhancement of the CD24–CD44+ population by jagged1 treatment was reversed upon reexpression of miR-205 (Supplemental Figure 1, C and D). To further delineate the role of miR-205 in the regulation of stem cell properties, we stably knocked down miR-205 using lentivirus-delivered shRNA (sh–miR-205) in MCF12A mammary epithelial cells. Suppression of miR-205 converted the epithelial phenotype to the mesenchymal phenotype, which was accompanied by a significant increase in expression levels of mesenchymal markers ZEB1 and N-cadherin and a decrease in the expression of the epithelial marker E-cadherin (Figure 1, F–H). Furthermore, KD of miR-205 elevated the CD24–CD44+ mammary stem cell population as well as the mammosphere formation (Figure 1, I and J) without inducing significant change in cell death or cell-cycle regulation (Supplemental Figure 1E). Together, these data suggest that miR-205 is critical for maintenance of the differentiated epithelial phenotype. And repressed miR-205 expression, in response to microenvironmental stimuli such as jagged1, is able to induce EMT and promote the stemness phenotype in both mammary epithelial cells and breast cancer cells.
NOTCH2 is a bona fide miR-205 target involved in stemness and cancer. To inquire about the direct target or targets of miR-205 that may be involved in the regulation of stem cell properties and tumorigenesis, we collected miR-205 putative targets that were overlapping among 3 miRNA-target prediction databases (DIANSmT, miRand, Targetscan), followed by DAVID Annotation Bioinformatics Database analysis to generate 16 candidates that had functional annotations involved in cell differentiation as well as in cancer (Figure 2A). Real-time PCR analysis was used to screen these candidates in the miR-205 KD cells compared with the control MCF12A cells. Among the 16 targets, only NOTCH2 and NOTCH4 showed significant elevation in response to the KD of miR-205 at the mRNA level (Supplemental Figure 2A). Immunoblot analysis revealed that the protein expression of NOTCH2 (full length and NICD), but not NOTCH4, was consistently upregulated in miR-205 KD cells and downregulated in cells ectopically expressing miR-205 (Supplemental Figure 2, B and C). To further validate that NOTCH2 is transcriptionally regulated by miR-205 through interaction of the complementary 3′ UTR region (Figure 2B), a luciferase reporter linked with 3′ UTR of NOTCH2 was used. Indeed, expression of miR-205 significantly decreased NOTCH2 luciferase activity, whereas a reporter mutated in NOTCH2–3′ UTR failed to respond to miR-205 expression (Figure 2C). Consistent with this observation, KD of miR-205 upregulated the protein level of NOTCH2 while ectopically expressing miR-205 downregulated NOTCH2 in MCF12A cells (Figure 2D). Similarly, NOTCH2 protein was significantly reduced by reexpression of miR-205 in breast cancer cell lines (BT-549 and MDA-MB-231) (Figure 2E), where endogenous miR-205 was shown to be repressed (9). Both NOTCH2 mRNA and protein were found to be highly expressed in the CD24–CD44+ cell population, where miR-205 expression was reduced compared with the non-CD24–CD44+ cell population isolated from PT (Figure 2F). Furthermore, jagged1 treatment significantly downregulated miR-205 while increasing NOTCH2 expression (Figure 2G). Together, these data suggest that NOTCH2 is a miR-205 target that may be functionally involved in stem cell properties.
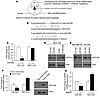 Figure 2
Figure 2NOTCH2 is a miR-205 target involved in stemness and cancer. (A) Schematic representation of in silico identification of the putative miR-205 targets involved in cell differentiation and cancer. (B) Diagram showing the putative miR-205–targeting seed sequence on NOTCH2–3′ UTR in humans and mice (underline indicates mutation of AAG to CCC in MUT-LUC). (C) Fold change in luciferase activity driven by the WT (WT-LUC) or mutant (MUT-LUC) NOTCH2–3′ UTR reporter under stable expression of miR-205 in BT549 breast cancer cells (n = 3, *P < 0.05). (D) Protein expression of NOTCH2 in MCF12A cells that stably expressed miR-205 or KD miR-205 (sh–miR-205). (E) Protein expression of NOTCH2 in breast cancer cells MDA-MB-231 and BT549 that stably expressed miR-205. (F) miR-205 and NOTCH2 expression levels in the CD44+CD24– stem cells versus the nonstem cells isolated from human PT (n = 3, *P < 0.05). (G) miR-205 and NOTCH2 mRNA expression under 40 μM jagged1 treatment in PT cells for 1 day (n = 3, *P < 0.05). Error bars denote mean ± SD.
NOTCH2 activation mediates induced EMT and stem cell properties caused by loss of miR-205. Previous gene expression microarray data demonstrated that NOTCH2 is one of the most upregulated genes in a cancer stem cell–like population (12) and its protein expression is significantly enhanced in the pancreatic cancer cells that acquire EMT (13). NOTCH activation has also been shown to target mouse mammary progenitor cells for expansion, leading to tumorigenesis (14). Based on the evidence, we asked whether NOTCH2 mediates miR-205–regulated EMT and stem cell properties. We found that KD of NOTCH2 to block the activation of NOTCH2 in miR-205 KD cells could significantly reduce the CD24–CD44+ cell population enhanced by loss of miR-205 (Figure 3, A and B). Similarly, in the aggressive breast cancer cell lines BT-549, MDA-MB-231, and PT, which exhibit repressed endogenous miR-205 expression and an enriched CD24–CD44+ cell population, reexpression of miR-205 resulted in a significantly reduced CD24–CD44+ cell population (Figure 3C), diminished sphere formation, and decreased ZEB1 expression; it also abolished mammary xenograft tumor formation in immunodeficient mice (Figure 3C, Supplemental Figure 3, A–D, and Table 2). The enhancement of the EMT and stemness phenotype caused by miR-205 suppression could be reversed by KD of NOTCH2 as well as by treatment of a γ-secretase inhibitor (GSI) (Figure 3, D and E, and Supplemental Figure 3, E and F). Together, these data suggest that NOTCH2 is a critical target that mediates miR-205–regulated stem cell properties in mammary epithelial cells and in breast cancer cells.
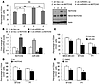 Figure 3
Figure 3miR-205 regulates stem cell properties through downregulation of NOTCH2. (A) The percentage of the CD24–CD44+ population (left) and NOTCH2 protein expression (right) in sh-Vec and sh–miR-205–MCF12A cells transfected with NOTCH2 siRNA (n = 3, *P < 0.05; **P < 0.01; sc, scrambled control siRNA). (B) Expression levels of miR-205 and NOTCH2 mRNA in sh-Vec and sh–miR-205–MCF12A cells transfected with NOTCH2 siRNA (n = 3, *P < 0.05; **P < 0.01). Error bars denote mean ± SD. (C) The percentage of CD24–CD44+ population in MB-MDA-231, BT549, and PT cells that stably expressed miR-205 (n = 3, *P < 0.05). (D) The percentage of CD24–CD44+ population in BT549 and PT cells transfected with NOTCH2 siRNA or (E) treated with 5 μM of GSI for 4 days (n = 3, *P < 0.05).
 Table 2
Table 2Transplanted mammary tumor formation in nude mice inoculated with the indicated number of PT cells expressing the control vector or miR-205
Loss of miR-205 leads to disrupted polarity and enhanced symmetric self renewal. Asymmetric division (AD) segregates cell fate determinants (e.g., Numb) exclusively into the daughter cells with specified cell fate, allowing only 1 daughter cell to maintain stem cell identity during stem cell division. However, studies have demonstrated that the stem cell pool can be expanded through self-renewing symmetric divisions (SDi), by which each stem cell produces 2 daughter cells with identical stem cell fate and proliferation potential at a time of wound repair or regeneration (15).
It is known that disrupted apical-basal cell polarity can lead to aberrant (unpolarized) localization of cell fate determinants, which in turn contributes to increased frequency of SDi of tumor stem cells and the expansion of the tumor stem cells to drive mammary tumor progression in vivo (16). Therefore, we asked whether miR-205 could affect cell polarity and consequent cell fate to regulate the stem cell population. Using a 3D Matrigel culture, we showed that loss of miR-205 in mammary epithelial cells led to enlarged, disorganized acinar formation with disrupted expression of the α6-integrin (basal polarity marker) and E-cadherin (epithelial marker) (Figure 4, A and B). Loss of miR-205 also significantly increased acinar luminal filling (as shown by the increased number of luminal nuclei vs. peripheral nuclei; Figure 4B), manifesting a cancerous phenotype.
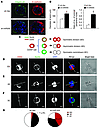 Figure 4
Figure 4Loss of miR-205 leads to disrupted polarity and enhanced symmetric self renewal. (A) Confocal fluorescence images showing acinar morphology of sh-Vec and sh–miR-205–MCF12A cells in 3D Matrigel culture. Cells were stained for basal polarity marker (α6-integrin, green) and epithelial marker (E-cadherin, red). Scale bars: 30 μm. (B) Bar graph showing altered acinar size and acinar luminal filling (as shown by the increased number of luminal nuclei vs. peripheral nuclei) in 3D-cultured sh-Vec and sh–miR-205–MCF12A cells. Error bars denote mean ± SD. (C) Schematic showing SDi, AD, and SC patterns. (D–F) Confocal fluorescence images showing CD44 (red) and NUMB (green) intracellular distribution during the first cell division of the CD24–CD44hi population isolated from MCF12A cells expressing sh-Vec and sh–miR-205 and cultured in suspension with the blebbistatin treatment. Scale bars: 20 μm. (G) Pie charts showing the percentage of the SDi versus AD patterns (n = 3, 70–110 cells were counted per sample, *P < 0.05).
Next, we examined the effect of miR-205 in the polarity of stem cell division by analyzing the intracellular distribution of a cell fate determinant NUMB immediately after mitosis. A stem cell–enriched CD24–CD44hi population isolated from MCF12A cells stably expressing sh–miR-205 or the control vector was treated with blebbistatin, a small molecule that arrests cytokinesis to form a binucleated cell and that was used for studying symmetric versus asymmetric cell division (ref. 16 and Figure 4C). We found that CD44 expression was lost in up to 48% of the dividing vector control cells that showed a uniform distribution of NUMB around the cell cortex of the binucleated cell (Figure 4, C, F, and G), suggesting that both of the progeny cells may share a symmetric cell fate in losing the stem cell identity (symmetric commitment [SC]). Another 45% of the control cells showed asymmetric distribution of CD44 and NUMB (AD), where NUMB was only localized to the CD44-negative progeny, but not to the stem cell–like progeny expressing a high level of CD44 (Figure 4, C, E, and G). Less than 7% of the control cells showed symmetric expression of CD44 along with a uniform distribution of NUMB in the binucleated cell, manifesting an SDi phenotype. Interestingly, this SDi phenotype was substantially enhanced upon suppression of miR-205 (7% to 45%; Figure 4, C, D, and G). Consistently, reexpression of miR-205 in the CD24–CD44hi population isolated from the human PT led to a significantly reduced frequency of SDi compared with the control cells (62% vs. 14%; Supplemental Figure 4, A and B). Together, these data suggest that loss of miR-205 may contribute to expansion of the stem cell/tumor stem cell population through promoting self-renewing SDi and that restoration of miR-205 could effectively diminish the cancer stemness by manipulating the stem cell fate.
Loss of miR-205 promotes SDi and self renewal through coordinated upregulation of ZEB1 and NOTCH2. To delineate the roles of NOTCH2 and ZEB1 in mediating miR-205–regulated stem cell division and cell fate, we knocked down NOTCH2 or ZEB1 expression in sh–miR-205–MCF12A cells using siRNA. Again, by analyzing the intracellular distribution of NUMB in the CD44hi population, we found that suppression of NOTCH2 not only abolished the enhancement of the SDi phenotype caused by KD of miR-205, but promoted a predominant SC pattern (Figure 5, A and B). Moreover, KD of miR-205 significantly upregulated expression levels of a cohort of stemness-related genes, including BMI1, MYC, NANOG, KLF4, OCT4, SOX2, and SOX9, and the enhancement of SOX9, NANOG, and KLF4 could be reversed upon suppression of NOTCH2 (Figure 5D). Consistently, enforced NOTCH2 expression reversed the effects of expression of miR-205, as shown by an enhancement of the SDi phenotype, accompanied by a substantial reduction of SC and increased expression of SOX9 and NANOG (Supplemental Figure 4, C and E). Together, these data suggest that NOTCH2 may play a role in maintaining the stemness state of the stem cell population promoted by suppression of miR-205.
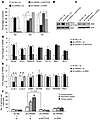 Figure 5
Figure 5Loss of miR-205 promotes self-renewing, symmetrically dividing stem cell population through coordinated activation of NOTCH2 and ZEB1. (A) The percentage of the symmetric and asymmetric cell division patterns in the isolated CD44hi-MCF12A cells expressing control sh-Vec and sh–miR-205 along with NOTCH2 or ZEB1 siRNA (n = 3, 70–110 cells were counted per sample, *P < 0.05). Protein expression levels of (B) NOTCH2 and (C) ZEB1 in the isolated CD44hi population expressing the indicated constructs. (D) mRNA expression of stemness-related genes in MCF12A cells expressing sh-Vec, sh–miR-205, or sh–miR-205 with NOTCH2 siRNA (n = 3, *P < 0.05). (E) mRNA expression of polarity genes in MCF12A cells expressing sh-Vec, sh–miR-205, or sh–miR-205 with ZEB1 siRNA (n = 3, *P < 0.05). (F) Number of forming spheres per 1,000 cells generated from MCF12A cells expressing sh-Vec, sh–miR-205, sh–miR-205 with NOTCH2 siRNA, or sh–miR-205 with ZEB1 siRNA for 3 serial passages (n = 3, *P < 0.05 compared with primary sphere in each group). Error bars denote mean ± SD.
Furthermore, it was demonstrated that loss of polarity protein expression can disrupt asymmetric distribution of Numb and induce Drosophila neuroblasts to divide symmetrically, leading to transplantable tumors that resemble mammalian cancers (17). Interestingly, in contrast with NOTCH2, we found that suppression of ZEB1 in the miR-205 KD stem cells switched the SDi phenotype to a predominant AD phenotype (Figure 5, A and C). Thus, we hypothesized that ZEB1, serving as a transcription repressor, may play a role in modulating asymmetric-symmetric cell division through transcriptional regulation of polarity protein expression. Indeed, our results showed that, among the polarity proteins that have been implicated in breast cancer (CRB3, DLG, LLGL1, LLGL2, SCRIB, PARD6A, PKCI) (18), KD of miR-205 significantly downregulated the gene expression of LLGL1 and LLGL2 (lethal 2 giant larvae homolog), which was rescued by a further KD of ZEB1 (Figure 5E). Consistently, enforced ZEB1 expression reversed the effects of expression of miR-205. as shown by an enhancement of the SDi phenotype, accompanied by a substantial reduction of AD and decreased expression of LLGL1 and LLGL2 (Supplemental Figure 4, D and E).
The promoter analysis further revealed 2 putative ZEB1-binding elements located in the LLGL1 promoter (A1, B1) and another 2 elements located in the LLGL2 promoter (A2, B2) within a 2-kb region upstream of the transcription start site (Supplemental Figure 5A). ChIP analysis showed that ZEB1 was significantly associated with A1 and B2 elements of LLGL1 and LLGL2, respectively (Supplemental Figure 5B), and the luciferase result validated that ZEB1 transcriptionally repressed LLGL1 and LLGL2 through binding to the specific promoter regions (Supplemental Figure 5C). KD of LLGL1 and LLGL2 reduced the AD stem cell population, but markedly increased the SDi stem cells (Supplemental Figure 5D), recapitulating the effects of KD of miR-205.
Furthermore, by serial mammosphere formation analysis, we showed that loss of miR-205 serially increased the number of mammospheres at each passage for 3 passages, potentially by promoting the self-renewing SDi. On the one hand, KD of NOTCH2 in miR-205–deficient cells substantially diminished sphere formation at each passage, suggesting an impairment of stemness maintenance, which leads to a significant reduction in the number of the primary sphere formation (P = 0.01) and continuing loss of stem cell population at each passage (Figure 5F). On the other hand, KD of ZEB1 in the miR-205–deficient cells led to a modest decrease in the number of the primary sphere formation (P = 0.05 as compared with the sh–miR-205 primary sphere), which was around the same number of secondary and tertiary spheres, suggesting an impairment in the SDi mechanism that failed to expand the stem cell pool; however, a constant stem cell population can still be maintained through AD at each passage (Figure 5F). Together, these data suggest that miR-205 plays a role in the polarity of stem cell division and cell fate through coordinated regulation of ZEB1 and NOTCH2.
Loss of miR-205 results in development of mammary premalignant and malignant lesions with increased EMT and stem cell properties in vivo. To provide physiological evidence for miR-205 regulation of cancer stemness in vivo, we established mammary-specific miR-205 KD mice by delivering concentrated lentiviruses that stably expressed miR-205–shRNA or control shRNA (miR-205 sequence is identical in humans and mice) into the mammary epithelia of WT BALB/c female mice (n = 10 mice/each group) via a well-established intramammary ductal injection technique (19–21). In contrast to the characteristic ductal outgrowth of the whole-mount mammary glands transduced with the control virus, more than 90% of the miR-205 KD mammary glands exhibited increased ductal length, extensive side branching, enlarged alveolar buds (Figure 6, A and B), and development of hyperplastic lesions with substantial fibrosis in 3 weeks (2% vs. 45%, Figure 6C; 3% vs. 36%, Figure 6D; collagen deposition as a marker of fibrosis is stained light blue in Figure 6D). Interestingly, fibrosis is known to be associated with breast cancer development, and accumulated evidence indicates that fibrogenic cells, such as myofibroblasts, can be induced from epithelial cells via EMT (22). Similarly to the in vitro data, KD of miR-205 elevated the expression of NOTCH2 and ZEB1 along with a significant reduction in E-cadherin levels compared with the control mammary tissues (Figure 6E). Furthermore, loss of miR-205 led to an increased mouse mammary stem cell population marked by lin–CD24+CD29hi cells (Figure 6F). Notably, 3 months after virus introduction, approximately 60% of the miR-205 KD mammary glands started to display focal mammary carcinoma (Figure 6G; n = 10 mice/each group, P < 0.03), whereas no hyperplasia or tumor was observed in the control mice. Consistently, the tumor sphere formation was significantly enhanced by loss of miR-205, and the enhancement could be reversed by GSI treatment (Figure 6H). To further understand whether GSI treatment could be therapeutically effective for the premalignant and malignant mammary lesions caused by loss of miR-205, we treated 2 groups of mammary-specific miR-205 KD mice along with the control mice with GSI (5 mg/kg) at 3 weeks (n = 5 mice) after virus introduction and also at 3 months (n = 5 mice) after virus introduction, when the miR-205 KD mammary glands started to display hyperplastic lesions and mammary carcinoma, respectively. Consistent with the in vitro observations, GSI treatment was able to revert both the premalignant and malignant phenotypes of the miR-205 KD mammary glands to a phenotype similar to the control glands, accompanied by diminished mammary carcinoma (Supplemental Figure 6, A, C, and D), decreased mammary stem cell population, and also reduced tumor sphere formation (Supplemental Figure 6, B and E). Together, these data suggest that loss of miR-205 results in enhanced EMT and stemness phenotype in mouse mammary glands, potentially contributing to the occurrence of premalignant and malignant mammary lesions. Suppression of NOTCH signaling by GSI treatment can be an effective therapeutic agent for treating miR-205–deficient breast lesions, potentially by eliminating the cancer stemness.
 Figure 6
Figure 6Loss of miR-205 in murine mammary glands promotes EMT and stemness phenotype, resulting in mammary tumorigenesis. (A) Upper panels: whole-mount images of the control (left) and miR-205 KD (right) mammary glands. Scale bars: 5 mm. Lower panels: magnification of the corresponding regions (enclosed) in the glands. Scale bars: 200 μm. (B) Left, bar graph showing the length of the ductal tree measured from the lymph node center to the end of ductal tree (boxed area in A). Right, bar graph showing the number of the branching points per 1 mm (n = 10/group, *P < 0.05). Images showing (C) H&E staining (left) and quantification of fibrosis foci (right), (D) Masson’s trichrome staining (left) and quantification of trichrome-blue area (right) of the control and miR-205 KD mammary tissues. Scale bars: 50 μm. *P < 0.05. Collagen is stained light blue, nuclei are stained dark purple, and cytoplasm and muscle fibers are stained red. (E) Bar graph showing miR-205/mRNA expression and (F) the percentage of lin–CD24+CD29hi cell population from the indicated mammary glands. (G) Representative miR-205/NOTCH2/ZEB1 staining in control tissue and miR-205 KD mammary tumor sections. Scale bars: 20 μm. (H) The number of primary spheres per 1,000 cells isolated from the control tissues and miR-205 KD mammary tumors. miR-205 KD tumor spheres were further treated with 5 μM DMSO or GSI for 4 days. Scale bars: 100 μm. n = 10/group, *P < 0.05. Error bars denote mean ± SD.
Loss of miR-205 is positively correlated with enhanced expression levels of NOTCH2 and ZEB1 in aggressive human breast tumors. To further strengthen the pathological correlation of miR-205–regulated signaling cascade in human breast cancer, we performed a correlation analysis of miR-205 expression with protein levels of ZEB1 as well as the newly identified target NOTCH2 in human breast cancer tissue microarrays consisting of 98 breast tumor samples. We found that, consistently, loss of miR-205 was positively correlated with an enhanced level of NOTCH2 (Figure 7A and Table 3; n = 98, P = 0.003). Ninety-four cases of these breast cancer samples were further grouped into 3 breast cancer subtypes: luminal (A+B), HER2+, and TNBC (4 cases did not belong to any of the subtypes). Our data revealed a very strong correlation between low/negative miR-205 expression (P = 0.0003) and high NOTCH2 expression (P = 0.0001) with the TNBC subtype (Figure 7B and Table 4). In contrast, high miR-205 expression and low/negative NOTCH2 expression were positively correlated with the luminal subtype; no correlation was found to HER2+ breast cancer (Figure 7B and Table 4). Furthermore, miR-205 expression (P = 0.00001) was most significantly reduced in the poorly differentiated high-grade tumors where JAG1 (P = 0.003), ZEB1 (P = 0.01), and NOTCH2 (P = 0.03) were also most highly expressed (Figure 7B and Table 5). Interestingly, it has been shown that high-grade tumors and TNBCs are often enriched in abundant tumor stem cell populations (2), which is consistent with our findings that dysregulated miR-205 signaling not only functionally links to tumor stem cell traits, but is highly correlated with aggressiveness and poor differentiation in breast cancer.
 Figure 7
Figure 7Loss of miR-205 is positively correlated with enhanced expression levels of NOTCH2 and ZEB1 in aggressive human breast tumors. (A) Representative cases from 98 breast cancer specimens in tissue microarrays were analyzed by immunohistochemical staining (NOTCH2) and by in situ hybridization (miR-205). Scale bars: 100 μm. (B) Representative images showing a reverse correlation of miR-205 with NOTCH2, ZEB1, and JAG1 expression. Scale bars: 100 μm.
 Table 5
Table 5χ2 Analysis showing a reverse correlation between miR-205 and NOTCH2/ZEB1/JAG1 in low-grade tumors (grade I) vs. high-grade tumors (grades II–III)
-
Discussion
Emerging evidence shows that the machinery controlling cell polarity and AD has an evolutionarily conserved role in the regulation of tumorigenesis. A previous study proposed that in the WT neuroblasts, with Numb acting as a conserved inhibitor of Notch signaling, coordinated balance of the polarity proteins, including LGL, ensured asymmetric segregation of Numb into the progenitor cell with specified cell fate, where Numb sufficiently antagonized Notch signaling. However, in the Lgl mutant, redistribution of NUMB in the cortex of the symmetrically dividing neuroblasts potentially led to insufficient Numb to block the Notch activity in both of the progenitor cells and thereby dampened the proper differentiation (23). These findings together with our data suggest an intricate regulatory network that links cell polarity and cell fate decision, both of which could be controlled by miR-205 in order to maintain the homeostasis of the differentiated epithelial cell population versus the stem cell population in response to microenvironmental stimuli (e.g., jagged1), providing important implications for tissue regeneration and cancer (Figure 8).
 Figure 8
Figure 8miR-205 coordinately regulates NOTCH2 and ZEB1 to control stem cell self renewal and SD. (A and B) A proposed model illustrating that jagged1-NOTCH signaling downregulates miR-205 and loss of miR-205 promotes self renewal and SD through constitutive activation of NOTCH2 and ZEB1, leading to expansion of stem cell and tumor stem cell populations.
Despite the report by Greene et al. (24) demonstrating that miR-205 expression increases the mouse mammary stem/progenitor population, the study was mostly based on examination of the Sca1+ cells isolated from 1 mouse mammary epithelial cell line, COMMA-DβGeo. The results actually contradicted those of other studies (25, 26), which showed that Sca1+ cells from the primary mouse mammary epithelial cells could not retain stem cell/progenitor activity, nor did they enrich the CD29hiCD24+ mouse mammary stem cell population. Additionally, Sca1+ cells are luminal (estrogen receptor positive) (27), which may represent a different population from the cells that harbor EMT and CD24–CD44+ stem cell phenotype (basal like) as shown in our study (28).
Furthermore, similar to our observation with miR-205, it was shown that p53 regulated the polarity of cell division in mouse mammary stem cells, and loss of p53 led to uniform redistribution of NUMB around the cell cortex of the dividing stem cell, increased the replicative potential of the mammospheres, and enhanced the frequency of SDi (16). Although the mechanism by which p53 regulates SDi was not elucidated, we think that miR-205–regulated signaling may play a role in the process, as a recent study suggests that miR-205 was implicated as a direct transcription target of p53 (29).
A recent finding also illustrated a cooperative relationship between a stemness-related transcription factor SOX9 and an EMT-related transcription factor SLUG, which is required to convert mammary epithelial cells to mammary stem cells, as SLUG alone, although activating a robust EMT program, failed to induce mammary stem cells from differentiated luminal cells (30). It is well known that both SLUG and ZEB1 are critical transcription factors involved in the EMT process; more interestingly, our data revealed that SOX9 expression could be regulated by miR-205–NOTCH2 signaling (Figure 5A), and other reports have also shown that SOX9 is a NOTCH target gene (31). Together, these data would support the notion that loss of miR-205 coordinately activates the signaling cascades and targets (e.g., ZEB1, NOTCH2) that are essential for induction of EMT and reprogramming of the cell fate.
-
Methods
Cell culture and treatment. Immortal normal mammary epithelial cells, MCF12A, and breast cancer cell lines MDA-MB231 and BT549 were purchased from ATCC. Primary breast tumor cells (freshly isolated from grades II–III ductal carcinoma) were purchased from Promab Inc. MCF12A cells were grown in DMEM-F12 medium supplemented with 5% horse serum, EGF (20 ng/ml), insulin (10 μg/ml), cholera toxin (1 ng/ml), hydrocortisone, and gentamycin (Sigma-Aldrich). MDA-MB231 and BT549 cells were cultured with DMEM supplemented with 10% FBS, penicillin (50 U/ml), and streptomycin (50 U/ml). Primary HMECs were cultured in MEGM Mammary Epithelial Cell Growth Medium (Lonza Inc.). For the jagged1 treatment, cells were treated with 40 μM of jagged1 active peptide or control peptide (Anaspec) for 24 hours. For the NOTCH inhibitor GSI treatment, 5 μM γ-Secretase Inhibitor IX (DAPT, N-[N-(3,5-Difluorophenacetyl-L-alanyl)]-S-phenylglycine-t-Butyl Ester; EMD Millipore) or DMSO was used. miR-205 precursor, miR-205 antagomir, and negative control oligos (50 nmole) were purchased from Ambion. siRNAs (200 pmole) targeting NOTCH2 and ZEB1 were from Santa Cruz Biotechnology Inc. Transfection of miR-205 oligos as well as the above siRNAs was performed by Lipofectamine 2000 following the manufacturer’s instructions.
Plasmid construction. Lentiviral constructs of miR-205 precursor (pMIR-205), miR-205 inhibitor (pMIRZIP-205), and the control vectors (pCDH, pMIR, and pMIRZIP) were purchased from System Biosciences. The EGFP fragment was removed from the pMIRZIP vector so that pMIRZIP-205 only carried the puromycin resistance cassette for selection of stably infected cells. To generate pCDH–miR-205-puro expression plasmid, miR-205 precursor fragment from pMIR-205 was subcloned into pCDH-puro. NOTCH2–3′ UTR luciferase construct was purchased from Switchgear Genomics, and PCR-based site-directed mutagenesis (Stratagene) was used to generate the 3′ UTR mutant. All of these constructs were verified by DNA sequencing.
Generation of stably expressed and KD cell lines. Lentiviral infection was performed as described previously (8). Briefly, pCDH-puro, pCDH–miR-205–puro, pMIRZIP-puro, or pMIRZIP-205-puro was cotransfected with the third generation of lentiviral packaging plasmids: pMDLg-pRRE, PRSV-Rev, and PMD2G into 293T cells by Lipofectamine 2000 following the manufacturer’s instructions. At 24 and 48 hours after transfection, culture medium was harvested and subsequently incubated with target cells in the presence of polybrene (5 μg/ml) for 24 hours. At 48 hours after infection, infected cells were harvested for gene and protein expression analysis or selected with puromycin (2 μg/ml) for 2 weeks to establish stably infected cells. Expression plasmid of NOTCH2 N-terminal intracellular domain (NOCTH2-NICD) was provided by T.S. Yeh (National Yang-Ming University, Taipei, Taiwan). Expression plasmid of ZEB1, pReceiver-ZEB1, was purchased from Genecopoedia Inc. Cells with expression of ZEB1 or NOTCH2-NICD were selected with Hygromycin (100 μg/ml) or G418 (800 μg/ml) for 2 weeks to establish stably expressed cells, respectively.
Antibodies. The following antibodies were used for immunoblotting according to standard protocol: anti-ZEB1 (Cell Signaling, 1:1000), anti-NOTCH2 (Cell Signaling, 1:2500), anti-JAG1 (Abcam, 1:1000), anti–E-cadherin (Cell Signaling, 1:1000), and anti–N-cadherin ( Abcam, 1:1000).
Flow cytometry analysis. FITC-conjugated anti-CD44 (BD Biosciences), PE-conjugated anti-CD24 (BD Biosciences), or FITC-conjugated anti-CD29 (BD Biosciences) antibodies were used for FACS analysis as described previously (8). PI staining for cell-cycle analysis was performed according to the standard protocol. Data were analyzed by FCS3 express (Denovo Software).
ChIP assay. The sequence of MIR205 promoter was obtained from the UCSC Genome Database. Analysis of putative transcription factor binding sites on SLUG and the LLGL1/2 promoter was done by TRED (Cold Spring Harbor Laboratory) and MatInspector (Genomatix). ChIP experiments were modified from the EZ-CHIP (Upstate) protocol using HES1 antibody (Santa Cruz Biotechnology Inc.) and ZEB1 (Bethyl). Primer sequences used for HES1-ChIP were as follows: fragment A, TTTGGCTGGTTTTTCAGTCC (forward), AAGCACATGCACGATCAGAG (reverse); fragment B, TACCCATCTTGGAGGGTACG (forward); TCCTGCTCTGCCTTCTTGAT (reverse).
Real-time PCR. Total RNAs were extracted from cells by using the RNeasy Kit (QIAGEN). RNAs were reverse transcribed by using the Superscript II Kit (Invitrogen). Results were analyzed by the iCycler (Bio-Rad), and the quantification of RNA levels was normalized to GAPDH as follows: CT (difference of cycling threshold) = CT (target) – CT (control). Higher CT values indicate relatively lower expression of RNA levels.
Immunocytochemistry and in situ hybridization. Breast cancer tissue microarrays and tissue sections were purchased from Pantomics and immunostained with anti-ZEB1 (Bethyl Laboratories, 1:100), anti-JAG1 (Cell Signaling, 1:500 for soluble JAG1 in Figure 1B; Abcam 1:100 for full-length JAG1 in Figure 7B), and anti-NOTCH2 (Cell Signaling, 1:2000). The histological grading and pathological annotation (tumor grade and subtype) were provided by the pathologists at Pantomics. Detection of the antibody signal was performed with the LSAB2 System-HRP (K0672; Dako) and the liquid DAB+ Substrate Chromogen System (K3468; Dako) according to the manufacturer’s instructions. In situ hybridization was performed using the miR-205 probe from Exiqon (miRCURY LNA Detection Probe; 5′-DIG-labeled). Detection of the probe was carried out using digoxigenin antibody (Abcam ab420, 1:200), LSAB2 System-HRP (K0672, Dako), and the liquid DAB+ Substrate Chromogen System (K3468, Dako) according to the manufacturer’s instructions. Cell nuclei were counterstained with Mayer’s hematoxylin stain. The correlation between miR-205 and ZEB1 and NOTCH2 was analyzed using the χ2 test. Trichrome staining was performed as described previously (32). Quantitation of the trichrome-positive area was performed by ImageJ analysis (http://imagej.nih.gov/ij/).
3D Matrigel culture. Cells were embedded between 2 layers of Matrigel (BD Biosciences) on 24-well plates. Wells were precoated with Matrigel (120 μl/well) to allow polymerization at 37°C for 15 minutes. Cells were then seeded at 20,000 cells/well density. After attachment (30 minutes at 37°C), cells were covered with a second layer of Matrigel/culture medium (1:19, 5%) to polymerize overnight at 37°C. Cells were incubated for 16 days, and the medium was replenished every 2 days. At the end of incubation, cells were fixed and subjected to immunofluorescence analysis.
Immunofluorescence staining. Cells were fixed using formalin for 20 minutes at room temperature (RT). Next, cells were permeabilized with 0.5% Triton X-100 in PBS for 5 minutes at RT and washed 3 times with 100 mM glycine at RT. Fixed cells were blocked for 1.5 hours with 10% goat serum. Cells were incubated overnight at 4°C with primary antibodies. The primary antibodies used were as follows: rat anti–α6-integrin (Millipore; 1:200 in Dako antibody diluent buffer) and rabbit anti–E-cadherin (Santa Cruz Biotechnology Inc., 1:100 in Dako antibody diluent buffer). Cells were incubated with secondary antibody for 1 hour, followed by 3 washes at RT. Secondary antibodies were as follows: FITC goat anti-rat and rhodamine goat anti-rabbit (Jackson ImmunoResearch). Cell nuclei were counterstained and mounted with Prolong Gold Antifade Reagent with DAPI (Molecular Probes) overnight at RT.
Cell division. The indicated CD24–CD44hi cells were treated with 25 mM blebbistatin for 48 hours in suspension culture, fixed for 10 minutes in 10% neutral buffered formalin, permeabilized with 0.1% Triton X-100 and 3% BSA, and costained with anti-Numb (Millipore, 1:500 dilution), followed by anti-mouse FITC (Jackson Laboratories) antibodies, and anti-CD44 (Thermo Scientific, 1:200 dilution), followed by anti-rabbit rhodamine. Cell nuclei were counterstained and mounted with Prolong Gold Antifade Reagent with DAPI (Molecular Probes) overnight at RT.
Confocal microscopy. Confocal laser scanning microscopy experiments were conducted using a Bio-Rad MRC-1024 laser-scanning confocal attachment mounted on a Nikon Diaphot 300 inverted microscope. More than 10 fields were imaged and analyzed per sample.
Intramammary ductal injection. Lentivirus was generated as indicated above and concentrated by Lenti-X Concentrator (Clontech). Virus titers were determined by serial dilution and virus-expressed EGFP expression. Equal titered viruses (1 × 107 TU/ml) expressing the control vector and sh–miR-205 were used to infect mammary glands of 4-month-old female BALB/c mice of 15-day lactation (n = 10 mice/group, glands on the left side were treated by sh–miR-205 viruses, and glands on the right side was treated by the sh-vector; 25 μl for each gland) using intramammary ductal injection through the nipples as described previously (19–21). To reduce intraductal pressure, the 15-day-old pups were separated from their mothers for 4 hours and then allowed to suckle for 30 to 45 minutes to deplete most of the milk in the mammary glands right before injection. To minimize potential mechanical damage, the injection rate was maintained at approximately 20 μl per 30 seconds (21).
Around 90% of primary mammary epithelial cells isolated from the mouse glands were successfully infected with the viruses (quantitated by EGFP expression using flow cytometry). To validate virus infection rate by intraductal injection, a total of 87 (control) and 55 (sh–miR-205) glands were examined in 10 nonconsecutive tissue sections from each injected gland (n = 5 glands/group); 76 ± 2 (control) and 51 ± 4 (sh–miR-205) glands showed EGFP staining in 3 independent infection experiments (P < 0.05). No positive staining was observed in the mock virus–infected glands. Approximately 90% of the glands were successfully KD for miR-205, as quantified by immunohistochemistry (IHC) staining of EGFP and miR-205. Three weeks after injection, mammary tissues from the euthanized mice were collected for whole-mount, IHC, H&E and Masson’s trichrome staining. Cells were isolated from the tissues for extraction of RNA, flow cytometry staining, and culture of primary spheres. For in vivo treatment, mice were treated with 5 mg/kg GSI (γ-secretase Inhibitor IX; Calbiochem) or DMSO via intraperitoneal injection every other day for 2 weeks at the indicated time points. The treatment regimen was designed based on previous studies (33–35).
Mammary gland whole mount. Dissected mouse mammary glands were placed into tissue cassettes and fixed overnight in 10% neutral buffered formalin. Mammary glands for whole mount were defatted in acetone, hydrated in ethanol, and stained with Mayer’s hematoxylin stain. Tissues were then destained and cleared in xylene and stored in methyl salicylate until photographed. To clearly exhibit the mammary ductal network, carmine alum dye (07070, Stem Cell Technologies) was used for staining of mammary glands according to the manufacturer’s instructions.
Mammary xenograft tumor. Mammary fat pads of female nude mice were inoculated with the indicated number of cells from human PT for a total volume of 100 μl per injection site. After 12 weeks, all of the tumors were calculated and animals were sacrificed.
Statistics. All data are presented as mean ± SD. Statistical calculations were performed with Microsoft Excel analysis tools. Differences between individual groups were analyzed by paired, 2-tailed t test. P < 0.05 was considered statistically significant. Genome-wide microRNA PCR array analysis was performed according to the manufacturer’s instructions (SABiosciences). χ2 Analyses for tumor samples were performed as described previously (8).
Accession numbers. All original microarray data were deposited in the NCBI’s Gene Expression Omnibus database (GSE56801).
Study approval. All experimental procedures were conducted with the approval of the Purdue Institutional Review Board and the Purdue Animal Care and Use Committee. All surgical procedures and animal manipulations were performed under the regulations of the Purdue Animal Care and Use Committee.
-
Acknowledgments
We thank Yava L. Jones-Hall (Department of Comparative Pathobiology, Purdue University College of Veterinary Medicine) for her assistance in histological examination of the mouse mammary tissue samples. This study was supported by a Showalter Research Scholar grant (206793 to C-J. Chang); P30 CA023168 to the Purdue University Center for Cancer Research in support of the use of facilities; an American Cancer Society Institutional Research grant to the Purdue University Center for Cancer Research (IRG 58-006-53) in support of the studies led by C.J. Chang and J.Y. Yang; the CPRIT Graduate Scholar Training Program at The University of Texas MD Anderson Cancer Center (to H.W. Ko); the National Breast Cancer Foundation Inc.; the China Medical University Hospital Cancer Research Center of Excellence (MOHW103-TD-B-111-03), and the Program for Stem Cell and Regenerative Medicine Frontier Research (NSC 102-2321-B-039-001 to M.C. Hung).
Address correspondence to: Chun-Ju Chang, Department of Basic Medical Sciences, 625 Harrison St., Purdue University, West Lafayette, Indiana 47907, USA. Phone: 765.494.2648; Fax: 765.494.0781; E-mail: chunjuchang@purdue.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Reference information: J Clin Invest. 2014;124(7):3093–3106. doi:10.1172/JCI73351.
Da Wang’s present address is: University Medical Center Groningen, Groningen, The Netherlands.
-
References
- Reya T, Morrison SJ, Clarke MF, Weissman IL. Stem cells, cancer, and cancer stem cells. Nature. 2001;414(6859):105–111.
- Pece S, et al. Biological and molecular heterogeneity of breast cancers correlates with their cancer stem cell content. Cell. 2010;140(1):62–73.
- Visvader JE, Lindeman GJ. Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat Rev Cancer. 2008;8(10):755–768.
- Wang Z, Li Y, Banerjee S, Sarkar FH. Emerging role of Notch in stem cells and cancer. Cancer Lett. 2009;279(1):8–12.
- Boulter L, et al. Macrophage-derived Wnt opposes Notch signaling to specify hepatic progenitor cell fate in chronic liver disease. Nat Med. 2012;18(4):572–579.
- Nyfeler Y, et al. Jagged1 signals in the postnatal subventricular zone are required for neural stem cell self-renewal. EMBO J. 2005;24(19):3504–3515.
- Lu J, et al. Endothelial cells promote the colorectal cancer stem cell phenotype through a soluble form of Jagged-1. Cancer Cell. 2013;23(2):171–185.
- Chang CJ, et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat Cell Biol. 2011;13(3):317–323.
- Sempere LF, et al. Altered MicroRNA expression confined to specific epithelial cell subpopulations in breast cancer. Cancer Res. 2007;67(24):11612–11620.
- Qin AY, et al. MiR-205 in cancer: an angel or a devil? Eur J Cell Biol. 2013;92(2):54–60.
- Gregory PA, et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol. 2008;10(5):593–601.
- Charafe-Jauffret E, et al. Breast cancer cell lines contain functional cancer stem cells with metastatic capacity and a distinct molecular signature. Cancer Res. 2009;69(4):1302–1313.
- Mazur PK, et al. Notch2 is required for progression of pancreatic intraepithelial neoplasia and development of pancreatic ductal adenocarcinoma. Proc Natl Acad Sci U S A. 2010;107(30):13438–13443.
- Bouras T, et al. Notch signaling regulates mammary stem cell function and luminal cell-fate commitment. Cell Stem Cell. 2008;3(4):429–441.
- Morrison SJ, Kimble J. Asymmetric and symmetric stem-cell divisions in development and cancer. Nature. 2006;441(7097):1068–1074.
- Cicalese A, et al. The tumor suppressor p53 regulates polarity of self-renewing divisions in mammary stem cells. Cell. 2009;138(6):1083–1095.
- Caussinus E, Gonzalez C. Induction of tumor growth by altered stem-cell asymmetric division in Drosophila melanogaster. Nat Genet. 2005;37(10):1125–1129.
- Martin-Belmonte F, Perez-Moreno M. Epithelial cell polarity, stem cells and cancer. Nat Rev Cancer. 2012;12(1):23–38.View this article via: PubMed Google Scholar
- Siwko SK, et al. Lentivirus-mediated oncogene introduction into mammary cells in vivo induces tumors. Neoplasia. 2008;10(7):653–662.View this article via: PubMed Google Scholar
- Krause S, Brock A, Ingber DE. Intraductal injection for localized drug delivery to the mouse mammary gland. J Vis Exp. 2013;(80):50692.View this article via: PubMed Google Scholar
- Nguyen DD, Beeman N, Lewis M, Schaack J, Neville MC. Intraductal injection into the mouse mammary gland. In: Ip MM, Asch BB, eds. Methods In Mammary Gland Biology And Breast Cancer Research. New York, New York, USA: Kluwer Academic/Plenum Publishers; 2000:259–270.
- Radisky DC, Kenny PA, Bissell MJ. Fibrosis and cancer: do myofibroblasts come also from epithelial cells via EMT? J Cell Biochem. 2007;101(4):830–839.
- Haenfler JM, Kuang C, Lee CY. Cortical aPKC kinase activity distinguishes neural stem cells from progenitor cells by ensuring asymmetric segregation of Numb. Dev Biol. 2012;365(1):219–228.
- Greene SB, Gunaratne PH, Hammond SM, Rosen JM. A putative role for microRNA-205 in mammary epithelial cell progenitors. J Cell Sci. 2010;123(pt 4):606–618.View this article via: PubMed Google Scholar
- Shackleton M, et al. Generation of a functional mammary gland from a single stem cell. Nature. 2006;439(7072):84–88.
- Stingl J, et al. Purification and unique properties of mammary epithelial stem cells. Nature. 2006;439(7079):993–997.View this article via: PubMed Google Scholar
- Stingl J. Detection and analysis of mammary gland stem cells. J Pathol. 2009;217(2):229–241.
- Mani SA, et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell. 2008;133(4):704–715.
- Piovan C, et al. Oncosuppressive role of p53-induced miR-205 in triple negative breast cancer. Mol Oncol. 2012;6(4):458–472.
- Guo W, et al. Slug and Sox9 cooperatively determine the mammary stem cell state. Cell. 2012;148(5):1015–1028.
- Shih HP, et al. A Notch-dependent molecular circuitry initiates pancreatic endocrine and ductal cell differentiation. Development. 2012;139(14):2488–2499.
- Laffin B, et al. Loss of singleminded-2s in the mouse mammary gland induces an epithelial-mesenchymal transition associated with up-regulation of slug and matrix metalloprotease 2. Mol Cell Biol. 2008;28(6):1936–1946.
- Lee CW, Raskett CM, Prudovsky I, Altieri DC. Molecular dependence of estrogen receptor-negative breast cancer on a notch-survivin signaling axis. Cancer Res. 2008;68(13):5273–5281.
- Nefedova Y, Sullivan DM, Bolick SC, Dalton WS, Gabrilovich DI. Inhibition of Notch signaling induces apoptosis of myeloma cells and enhances sensitivity to chemotherapy. Blood. 2008;111(4):2220–2229.
- Rizzo P, et al. Cross-talk between notch and the estrogen receptor in breast cancer suggests novel therapeutic approaches. Cancer Res. 2008;68(13):5226–5235.
-
Version history
- Version 1 (June 9, 2014): No description
- Version 2 (July 1, 2014): No description



Copyright © 2024 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)



