Machine Learning Protocols And Methods
Machine Learning, a branch of Artificial Intelligence that gives computers the capability to learn without being explicitly programmed.
Machine Learning, a branch of Artificial Intelligence that gives computers the capability to learn without being explicitly programmed. It involves the development of algorithms that enable computers to learn from and make predictions or decisions based on data. Machine Learning is commonly used for tasks such as pattern recognition, data classification, and predictive modeling. It has applications in various fields, including image and speech recognition, medical diagnosis, financial forecasting, and recommendation systems. Machine Learning is essential for analyzing complex data, identifying patterns, and making informed decisions, making it a powerful tool in research, industry, and everyday technology.
Recently cited
-
ArticleNature Methods
Robust enumeration of cell subsets from tissue expression profiles
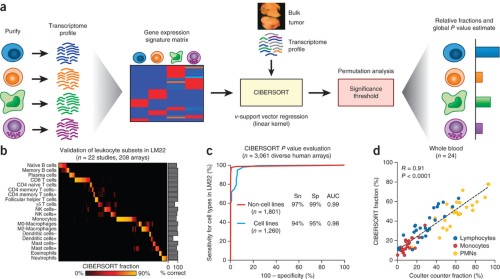
We introduce CIBERSORT, a method for characterizing cell composition of complex tissues from their gene expression profiles. When applied to enumeration of hematopoietic subsets in RNA mixtures from fresh, frozen and fixed tissues, including solid
…moreWe introduce CIBERSORT, a method for characterizing cell composition of complex tissues from their gene expression profiles. When applied to enumeration of hematopoietic subsets in RNA mixtures from fresh, frozen and fixed tissues, including solid tumors, CIBERSORT outperformed other methods with respect to noise, unknown mixture content and closely related cell types. CIBERSORT should enable large-scale analysis of RNA mixtures for cellular biomarkers and therapeutic targets (http://cibersort.stanford.edu/ ).
less -
PerspectiveNature Methods
The Perseus computational platform for comprehensive analysis of (prote)omics data
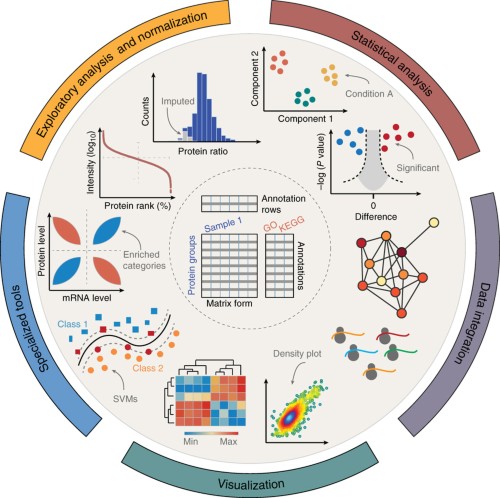
A main bottleneck in proteomics is the downstream biological analysis of highly multivariate quantitative protein abundance data generated using mass-spectrometry-based analysis. We developed the Perseus software platform
…moreA main bottleneck in proteomics is the downstream biological analysis of highly multivariate quantitative protein abundance data generated using mass-spectrometry-based analysis. We developed the Perseus software platform (http://www.perseus-framework.org ) to support biological and biomedical researchers in interpreting protein quantification, interaction and post-translational modification data. Perseus contains a comprehensive portfolio of statistical tools for high-dimensional omics data analysis covering normalization, pattern recognition, time-series analysis, cross-omics comparisons and multiple-hypothesis testing. A machine learning module supports the classification and validation of patient groups for diagnosis and prognosis, and it also detects predictive protein signatures. Central to Perseus is a user-friendly, interactive workflow environment that provides complete documentation of computational methods used in a publication. All activities in Perseus are realized as plugins, and users can extend the software by programming their own, which can be shared through a plugin store. We anticipate that Perseus's arsenal of algorithms and its intuitive usability will empower interdisciplinary analysis of complex large data sets.
less -
PerspectiveNature Methods
ilastik: interactive machine learning for (bio)image analysis
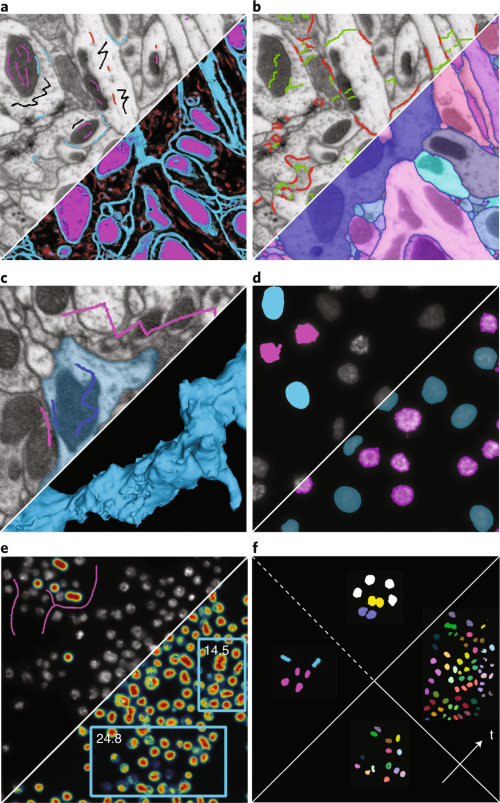
We present ilastik, an easy-to-use interactive tool that brings machine-learning-based (bio)image analysis to end users without substantial computational expertise. It contains pre-defined workflows for image segmentation, object classification,
…moreWe present ilastik, an easy-to-use interactive tool that brings machine-learning-based (bio)image analysis to end users without substantial computational expertise. It contains pre-defined workflows for image segmentation, object classification, counting and tracking. Users adapt the workflows to the problem at hand by interactively providing sparse training annotations for a nonlinear classifier. ilastik can process data in up to five dimensions (3D, time and number of channels). Its computational back end runs operations on-demand wherever possible, allowing for interactive prediction on data larger than RAM. Once the classifiers are trained, ilastik workflows can be applied to new data from the command line without further user interaction. We describe all ilastik workflows in detail, including three case studies and a discussion on the expected performance.
less
-
Brief CommunicationNature Methods
U-Net: deep learning for cell counting, detection, and morphometry
U-Net is a generic deep-learning solution for frequently occurring quantification tasks such as cell detection and shape measurements in biomedical image data. We present an ImageJ plugin that enables non-machine-learning experts to analyze their
…moreU-Net is a generic deep-learning solution for frequently occurring quantification tasks such as cell detection and shape measurements in biomedical image data. We present an ImageJ plugin that enables non-machine-learning experts to analyze their data with U-Net on either a local computer or a remote server/cloud service. The plugin comes with pretrained models for single-cell segmentation and allows for U-Net to be adapted to new tasks on the basis of a few annotated samples.
less -
Brief CommunicationNature Methods
NicheNet: modeling intercellular communication by linking ligands to target genes
Computational methods that model how gene expression of a cell is influenced by interacting cells are lacking. We present NicheNet (https://github.com/saeyslab/nichenetr), a method that predicts ligand–target links between interacting cells by
…moreComputational methods that model how gene expression of a cell is influenced by interacting cells are lacking. We present NicheNet (https://github.com/saeyslab/nichenetr), a method that predicts ligand–target links between interacting cells by combining their expression data with prior knowledge on signaling and gene regulatory networks. We applied NicheNet to tumor and immune cell microenvironment data and demonstrate that NicheNet can infer active ligands and their gene regulatory effects on interacting cells.
less -
This MonthNature Methods
Statistics versus machine learning
Danilo Bzdok , Naomi Altman , Martin KrzywinskiStatistics draws population inferences from a sample, and machine learning finds generalizable predictive patterns.
Statistics draws population inferences from a sample, and machine learning finds generalizable predictive patterns.
-
ArticleNature Methods
Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction
Cryogenic electron microscopy (cryo-EM) is widely used to study biological macromolecules that comprise regions with disorder, flexibility or partial occupancy. For example, membrane proteins are often kept in solution with detergent micelles and
…moreCryogenic electron microscopy (cryo-EM) is widely used to study biological macromolecules that comprise regions with disorder, flexibility or partial occupancy. For example, membrane proteins are often kept in solution with detergent micelles and lipid nanodiscs that are locally disordered. Such spatial variability negatively impacts computational three-dimensional (3D) reconstruction with existing iterative refinement algorithms that assume rigidity. We introduce non-uniform refinement, an algorithm based on cross-validation optimization, which automatically regularizes 3D density maps during refinement to account for spatial variability. Unlike common shift-invariant regularizers, non-uniform refinement systematically removes noise from disordered regions, while retaining signal useful for aligning particle images, yielding dramatically improved resolution and 3D map quality in many cases. We obtain high-resolution reconstructions for multiple membrane proteins as small as 100 kDa, demonstrating increased effectiveness of cryo-EM for this class of targets critical in structural biology and drug discovery. Non-uniform refinement is implemented in the cryoSPARC software package.
less -
ArticleNature Methods
Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs
Cryo-electron microscopy is a popular method for the determination of protein structures; however, identifying a sufficient number of particles for analysis can take months of manual effort. Current computational approaches find many false positives
…moreCryo-electron microscopy is a popular method for the determination of protein structures; however, identifying a sufficient number of particles for analysis can take months of manual effort. Current computational approaches find many false positives and require ad hoc postprocessing, especially for unusually shaped particles. To address these shortcomings, we develop Topaz, an efficient and accurate particle-picking pipeline using neural networks trained with a general-purpose positive-unlabeled learning method. This framework enables particle detection models to be trained with few sparsely labeled particles and no labeled negatives. Topaz retrieves many more real particles than conventional picking methods while maintaining low false-positive rates, is capable of picking challenging unusually shaped proteins (for example, small, non-globular and asymmetric particles), produces more representative particle sets and does not require post hoc curation. We demonstrate the performance of Topaz on two difficult datasets and three conventional datasets. Topaz is modular, standalone, free and open source (http://topaz.csail.mit.edu ).
less -
ArticleNature Methods
Large-scale automated synthesis of human functional neuroimaging data
The rapid growth of the literature on neuroimaging in humans has led to major advances in our understanding of human brain function but has also made it increasingly difficult to aggregate and synthesize neuroimaging findings. Here we describe and
…moreThe rapid growth of the literature on neuroimaging in humans has led to major advances in our understanding of human brain function but has also made it increasingly difficult to aggregate and synthesize neuroimaging findings. Here we describe and validate an automated brain-mapping framework that uses text-mining, meta-analysis and machine-learning techniques to generate a large database of mappings between neural and cognitive states. We show that our approach can be used to automatically conduct large-scale, high-quality neuroimaging meta-analyses, address long-standing inferential problems in the neuroimaging literature and support accurate 'decoding' of broad cognitive states from brain activity in both entire studies and individual human subjects. Collectively, our results have validated a powerful and generative framework for synthesizing human neuroimaging data on an unprecedented scale.
less -
ArticleNature Methods
Unified rational protein engineering with sequence-based deep representation learning
Rational protein engineering requires a holistic understanding of protein function. Here, we apply deep learning to unlabeled amino-acid sequences to distill the fundamental features of a protein into a statistical representation that is semantically
…moreRational protein engineering requires a holistic understanding of protein function. Here, we apply deep learning to unlabeled amino-acid sequences to distill the fundamental features of a protein into a statistical representation that is semantically rich and structurally, evolutionarily and biophysically grounded. We show that the simplest models built on top of this unified representation (UniRep) are broadly applicable and generalize to unseen regions of sequence space. Our data-driven approach predicts the stability of natural and de novo designed proteins, and the quantitative function of molecularly diverse mutants, competitively with the state-of-the-art methods. UniRep further enables two orders of magnitude efficiency improvement in a protein engineering task. UniRep is a versatile summary of fundamental protein features that can be applied across protein engineering informatics.
less
Recently published
-
ProtocolSpringer Protocols
Predicting Tumor Growth and Ligand Dependence from mRNA by Combining Machine Learning with Mechanistic Modeling
For successful treatment of cancer patients, it is crucial to identify subgroups that respond to certain types of targeted therapy. A key element for tumor growth is the abundance of receptors and binding of growth factors, which can be diminished
…moreFor successful treatment of cancer patients, it is crucial to identify subgroups that respond to certain types of targeted therapy. A key element for tumor growth is the abundance of receptors and binding of growth factors, which can be diminished via therapeutic antibodies. Here, a mechanistic signaling network model is linked to patient-specific ribonucleic acid sequencing data (RNAseq), enabling the prediction of individuals susceptible to a particular medication. The mechanistic model comprises multiple receptors and their dimerization, and is calibrated using time-resolved in-vitro data. Further, the model is combined with in-vitro cell viability measurements via a machine learning algorithm and ultimately applied to patient-derived data to predict ligand dependence of tumors. For this purpose, RNA sequencing data are exploited to constrain model parameters and generalize model response. Mathematical modeling of signal transduction is used as a mediator, performing a non-trivial transformation of initial protein expression levels and ligand conditions to cell-type specific response. Thereby, it allows for bridging the gap between studies of signal transduction on a short time scale and cell fate decisions in the long term, potentially aiding in drug development, patient stratification, and prediction of tumor response. This chapter is based on work previously published in Hass et al. (NPJ Syst Biol Appl 3(1):27, 2017) and Hass (Quantifying cell biology: mechanistic dynamic modeling of receptor crosstalk. PhD thesis, Albert-Ludwigs-Universität Freiburg, 2017).
less -
OverviewSpringer Protocols
Machine Learning Techniques for Development of Drugs Against Coronavirus Disease 2019 (COVID-19): A Case Study Protocol
Discovery of SARS-CoV-2 drug requires a fast track approach to achieve the effective and safe alternative. The present times call for an evolution in the process of drug discovery. Several factors contribute to high cost and long development times
…moreDiscovery of SARS-CoV-2 drug requires a fast track approach to achieve the effective and safe alternative. The present times call for an evolution in the process of drug discovery. Several factors contribute to high cost and long development times associated with new drugs. Finding a new molecular target is contingent upon a detailed understanding of the disease pathology, which often takes years of basic research. Integrating genetic and expression studies with Protein Interaction Network (PIN), and considering both functional and topological features of the resultant network may prove to be an effective target identification strategy. Further, apart from the existing computational tools to identify ligands, artificial intelligence approaches may now be used to increase the search space many folds, offering a faster method for screening. Artificial intelligence can be integrated with the existing drug discovery pipeline to enable rational target identification, prediction of an accurate 3D structure of the molecular target and screen large ligand libraries for putative modulators. The present chapter covers a detailed protocol to scan and validate the therapeutic targets for COVID-19, and screen the compounds for future in vitro or in vivo validation. The chapter covers target selection strategies, and application of artificial intelligence to identify drug–target interactions.
less -
OverviewSpringer Protocols
Deep Learning-Based Drug Screening for COVID-19 and Case Studies
Coronavirus infectious disease (COVID-19), caused by deadly severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has been declared as a pandemic by the World Health Organization. This disease has become the world’s worst infectious disease,
…moreCoronavirus infectious disease (COVID-19), caused by deadly severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has been declared as a pandemic by the World Health Organization. This disease has become the world’s worst infectious disease, killing about 1.05 million lives as of September 2020. The absence of vaccines and effective drugs is a key trouble responsible for the ineffective management of this pandemic. Considering this emergency situation, several trials have been made to identify repurposing on-market drugs known for their antiviral behavior. Several modern technologies such as deep machine learning are used to combat this deadly disease with faster prediction and greater accuracy. More interestingly, these studies have provided clues for the antiviral properties and are believed to help in effective control of this pandemic. The drugs identified by deep learning-based virtual screening will help in unraveling molecular mechanisms of therapeutic and antiviral properties and will pave the way for designing artificial drugs. Hence we focus in this chapter on the integrated applications of deep learning models as a pipeline for drug and vaccine discovery which has implications in therapeutic drug targeting for COVID-19.
less
-
ProtocolSpringer Protocols
ShapeMetrics: A 3D Cell Segmentation Pipeline for Single-Cell Spatial Morphometric Analysis
There is a growing need for single-cell level data analysis in correlation with the advancements of microscopy techniques. Morphology-based statistics gathered from individual cells are essential for detection and quantification of even subtle
…moreThere is a growing need for single-cell level data analysis in correlation with the advancements of microscopy techniques. Morphology-based statistics gathered from individual cells are essential for detection and quantification of even subtle changes within the complex tissues, yet the information available from high-resolution imaging is oftentimes sub-optimally utilized due to the lack of proper computational analysis software. Here we present ShapeMetrics, a 3D cell segmentation pipeline that we have developed to identify, analyze, and quantify single cells in an image. This MATLAB-based script enables users to extract morphological parameters, such as ellipticity, longest axis, cell elongation, or the ratio between cell volume and surface area. We have specifically invested in creating a user-friendly pipeline, aimed for biologists with a limited computational background. Our pipeline is presented with detailed stepwise instructions, starting from the establishment of machine learning–based prediction files of immuno-labeled cell membranes followed by the application of 3D cell segmentation and parameter extraction script, leading to the morphometric analysis and spatial visualization of cell clusters defined by their morphometric features.
less -
ProtocolSpringer Protocols
Online Tools and Antiviral Databases for the Development of Drugs Against Coronaviruses
The current crisis of coronavirus pandemic has created an urgent need for readily available scientific information to the researchers, students, professionals, and journalists. The scientific information for the research is costly, and most of the
…moreThe current crisis of coronavirus pandemic has created an urgent need for readily available scientific information to the researchers, students, professionals, and journalists. The scientific information for the research is costly, and most of the universities and research institutes cannot afford to subscribe to all the coronavirus-related journals/articles during this crisis time. In order to expedite the process of vaccine development and discovery of anti-COVID drugs, most of the pharmaceutical companies, research institutes and publishers are playing a key role and working on war footing to get either a vaccine or an anti-COVID drug as early as possible. The collective efforts are required from everyone in this testing time of the corona crisis. To provide our contribution to the scientific community, we have made here an attempt to give an overview of some of the tools and resources freely available that probably provide some insights in data mining and screening of novel lead molecules toward coronavirus. We have collected and compiled the information of open access online tools and antiviral databases essential for the discovery and development of corona vaccine and anti-COVID drugs. The open access tools include Open Educational Resources (OERs), Google cloud, online prediction server, web-based viewer, etc., while the antiviral databases include libraries of synthetic and untested compounds, antiviral drug databases, antiviral peptides, etc. The information presented in this chapter will help the researchers to use them directly in their projects of coronavirus drug discovery.
less -
ProtocolSpringer Protocols
A Lipidomics Approach to Determine the Role of Lipids and Its Crosstalk with Autophagy in Lung Cancer Metastasis
Non-small cell lung cancer (NSCLC) is among the most malignant tumors with high propensity for metastasis and is the leading cause of cancer-related death globally. Most patients present with regional and distant metastasis, associated with poor
…moreNon-small cell lung cancer (NSCLC) is among the most malignant tumors with high propensity for metastasis and is the leading cause of cancer-related death globally. Most patients present with regional and distant metastasis, associated with poor prognosis. Lipids may play an essential role in either activating or inhibiting detachment-induced apoptosis (anoikis), where the latter is a crucial mechanism to prevent metastasis, and it may have a cross-talk with autophagy. Autophagy has been shown to be induced in various human cancer metastasis, modulating tumor cell motility and invasion, cancer cell differentiation, resistance to anoikis, and epithelial to mesenchymal transition. Hence, it may play a crucial role in the transition of benign to malignant phenotypes, the core of metastasis initiation. Here, we provide a method we have established in our laboratory for detecting lipids in attached and detached non-small lung cancer cells and show how to analyze lipidomics data to find its correlation with autophagy-related pathways.
less -
OverviewSpringer Protocols
Machine Learning for Biological Design
We briefly present machine learning approaches for designing better biological experiments. These approaches build on machine learning predictors and provide additional tools to guide scientific discovery. There are two different kinds of objectives
…moreWe briefly present machine learning approaches for designing better biological experiments. These approaches build on machine learning predictors and provide additional tools to guide scientific discovery. There are two different kinds of objectives when designing better experiments: to improve the predictive model or to improve the experimental outcome. We survey five different approaches for adaptive experimental design that iteratively search the space of possible experiments while adapting to measured data. The approaches are Bayesian optimization, bandits, reinforcement learning, optimal experimental design, and active learning. These machine learning approaches have shown promise in various areas of biology, and we provide broad guidelines to the practitioner and links to further resources.
less -
ProtocolSpringer Protocols
Techniques for Developing Reliable Machine Learning Classifiers Applied to Understanding and Predicting Protein:Protein Interaction Hot Spots
With machine learning now transforming the sciences, successful prediction of biological structure or activity is mainly limited by the extent and quality of data available for training, the astute choice of features for prediction, and thorough
…moreWith machine learning now transforming the sciences, successful prediction of biological structure or activity is mainly limited by the extent and quality of data available for training, the astute choice of features for prediction, and thorough assessment of the robustness of prediction on a variety of new cases. In this chapter, we address these issues while developing and sharing protocols to build a robust dataset and rigorously compare several predictive classifiers using the open-source Python machine learning library, scikit-learn. We show how to evaluate whether enough data has been used for training and whether the classifier has been overfit to training data. The most telling experiment is 500-fold repartitioning of the training and test sets, followed by prediction, which gives a good indication of whether a classifier performs consistently well on different datasets. An intuitive method is used to quantify which features are most important for correct prediction. The resulting well-trained classifier, hotspotter, can robustly predict the small subset of amino acid residues on the surface of a protein that are energetically most important for binding a protein partner: the interaction hot spots. Hotspotter has been trained and tested here on a curated dataset assembled from 1046 non-redundant alanine scanning mutation sites with experimentally measured change in binding free energy values from 97 different protein complexes; this dataset is available to download. The accessible surface area of the wild-type residue at a given site and its degree of evolutionary conservation proved the most important features to identify hot spots. A variant classifier was trained and validated for proteins where only the amino acid sequence is available, augmented by secondary structure assignment. This version of hotspotter requiring fewer features is almost as robust as the structure-based classifier. Application to the ACE2 (angiotensin converting enzyme 2) receptor, which mediates COVID-19 virus entry into human cells, identified the critical hot spot triad of ACE2 residues at the center of the small interface with the CoV-2 spike protein. Hotspotter results can be used to guide the strategic design of protein interfaces and ligands and also to identify likely interfacial residues for protein:protein docking.
less -
OverviewSpringer Protocols
A Machine Learning Approach for Predicting Essentiality of Metabolic Genes
The identification of essential genes is a key challenge in systems and synthetic biology, particularly for engineering metabolic pathways that convert feedstocks into valuable products. Assessment of gene essentiality at a genome scale requires
…moreThe identification of essential genes is a key challenge in systems and synthetic biology, particularly for engineering metabolic pathways that convert feedstocks into valuable products. Assessment of gene essentiality at a genome scale requires large and costly growth assays of knockout strains. Here we describe a strategy to predict the essentiality of metabolic genes using binary classification algorithms. The approach combines elements from genome-scale metabolic models, directed graphs, and machine learning into a predictive model that can be trained on small knockout data. We demonstrate the efficacy of this approach using the most complete metabolic model of Escherichia coli and various machine learning algorithms for binary classification.
less -
ProtocolSpringer Protocols
Building a Binary Classification Machine-Learning Model: A Guide to Predicting Participation in a Lyme Disease Program at a Medical Institute
Kunal Garg ,
Liria
Mitzuko Fajardo-Yamamoto
,
Flor
Cecilia Rojas-Castro
,
Teo
Susnjak
,
Leona
Gilbert
,
Liria
Mitzuko Fajardo-Yamamoto
,
Flor
Cecilia Rojas-Castro
,
Teo
Susnjak
,
Leona
Gilbert
The field of data analysis, preparation, and machine learning is rapidly expanding, offering numerous libraries and resources for exploration. Researchers gain knowledge through various channels, but few resources provide a comprehensive framework
…moreThe field of data analysis, preparation, and machine learning is rapidly expanding, offering numerous libraries and resources for exploration. Researchers gain knowledge through various channels, but few resources provide a comprehensive framework for building machine-learningMachine-learning models. We present a step-by-step framework for constructing a robust Random Forest classification model to fill this gap. Using the trained model, we predict if individuals visiting Sanoviv Medical Institute between 2020 and 2023 participated in the Lyme disease program based on age, symptoms, blood count, and chemistry results. While not exhaustive, the methods in each step provide a valuable starting point for researchers, promoting an understanding of the fundamental approach to model creation. The framework encourages researchers to explore beyond the outlined techniques, fostering innovation and experimentation.
less
Review papers
-
OverviewSpringer Protocols
A Non-technical Introduction to Machine Learning
This chapter provides an introduction to machine learning for a non-technical readership. Machine learning is an approach to artificial intelligence. The chapter thus starts with a brief history of artificial intelligence in order to put machine
…moreThis chapter provides an introduction to machine learning for a non-technical readership. Machine learning is an approach to artificial intelligence. The chapter thus starts with a brief history of artificial intelligence in order to put machine learning into this broader scientific context. We then describe the main general concepts of machine learning. Readers with a background in computer science may skip this chapter.
less -
OverviewSpringer Protocols
Machine Learning and Brain Imaging for Psychiatric Disorders: New Perspectives
Psychiatric disorders include a broad panel of heterogeneous conditions. Among the most severe psychiatric diseases, in intensity and incidence, depression will affect 15–20% of the population in their lifetime, schizophrenia 0.7–1%, and bipolar
…morePsychiatric disorders include a broad panel of heterogeneous conditions. Among the most severe psychiatric diseases, in intensity and incidence, depression will affect 15–20% of the population in their lifetime, schizophrenia 0.7–1%, and bipolar disorder 1–2.5%. Today, the diagnosis is solely based on clinical evaluation, causing major issues since it is subjective and as different diseases can present similar symptoms. These limitations in diagnosis lead to limitations in the classification of psychiatric diseases and treatments. There is therefore a great need for new biomarkers, usable at an individual level. Among them, magnetic resonance imaging (MRI) allows to measure potential brain abnormalities in patients with psychiatric disorders. This creates datasets with high dimensionality and very subtle variations between healthy subjects and patients, making machine and statistical learning ideal tools to extract biomarkers from these data. Machine learning brings different tools that could be useful to tackle these issues. On the one hand, supervised learning can support automated classification between different psychiatric conditions. On the other hand, unsupervised learning could allow the identification of new homogeneous subgroups of patients, refining our understanding of the classification of these disorders. In this chapter, we will review current research applying machine learning tools to brain imaging in psychiatry, and we will discuss its interest, limitations, and future applications.
less
-
OverviewSpringer Protocols
Machine Learning for Parkinson’s Disease and Related Disorders
Parkinson’s disease is a complex heterogeneous neurodegenerative disorder characterized by the loss of dopamine neurons in the basal ganglia, resulting in many motor and non-motor symptoms. Although there is no cure to date, the dopamine replacement
…moreParkinson’s disease is a complex heterogeneous neurodegenerative disorder characterized by the loss of dopamine neurons in the basal ganglia, resulting in many motor and non-motor symptoms. Although there is no cure to date, the dopamine replacement therapy can improve motor symptoms and the quality of life of the patients. The cardinal symptoms of this disorder are tremor, bradykinesia, and rigidity, referred to as parkinsonism. Other related disorders, such as dementia with Lewy bodies, multiple system atrophy, and progressive supranuclear palsy, share similar motor symptoms although they have different pathophysiology and are less responsive to the dopamine replacement therapy. Machine learning can be of great utility to better understand Parkinson’s disease and related disorders and to improve patient care. Many challenges are still open, including early accurate diagnosis, differential diagnosis, better understanding of the pathologies, symptom detection and quantification, individual disease progression prediction, and personalized therapies. In this chapter, we review research works on Parkinson’s disease and related disorders using machine learning.
less
-
OverviewSpringer Protocols
Machine Learning for Cerebrovascular Disorders
Yannan Yu , David Yen-Ting ChenCerebrovascular disease refers to a group of conditions that affect blood flow and the blood vessels in the brain. It is one of the leading causes of mortality and disability worldwide, imposing a significant socioeconomic burden to society. Research
…moreCerebrovascular disease refers to a group of conditions that affect blood flow and the blood vessels in the brain. It is one of the leading causes of mortality and disability worldwide, imposing a significant socioeconomic burden to society. Research on cerebrovascular diseases has been rapidly progressing leading to improvement in the diagnosis and management of patients nowadays. Machine learning holds many promises for further improving clinical care of these disorders. In this chapter, we will briefly introduce general information regarding cerebrovascular disorders and summarize some of the most promising fields in which machine learning shall be valuable to improve research and patient care. More specifically, we will cover the following cerebrovascular disorders: stroke (both ischemic and hemorrhagic), cerebral microbleeds, cerebral vascular malformations, intracranial aneurysms, and cerebral small vessel disease (white matter hyperintensities, lacunes, perivascular spaces).
less -
OverviewSpringer Protocols
Machine Learning in Multiple Sclerosis
Multiple sclerosis (MS) is characterized by inflammatory activity and neurodegeneration, leading to the accumulation of damage to the central nervous system resulting in the accumulation of disability. MRI depicts an important part of the pathology
…moreMultiple sclerosis (MS) is characterized by inflammatory activity and neurodegeneration, leading to the accumulation of damage to the central nervous system resulting in the accumulation of disability. MRI depicts an important part of the pathology of this disease and therefore plays a key part in diagnosis and disease monitoring. Still, major challenges exist with regard to the differential diagnosis, adequate monitoring of disease progression, quantification of CNS damage, and prediction of disease progression. Machine learning techniques have been employed in an attempt to overcome these challenges. This chapter aims to give an overview of how machine learning techniques are employed in MS with applications for diagnostic classification, lesion segmentation, improved visualization of relevant brain pathology, characterization of neurodegeneration, and prognostic subtyping.
less -
OverviewSpringer Protocols
Classic Machine Learning Methods
Johann Faouzi , Olivier ColliotIn this chapter, we present the main classic machine learning methods. A large part of the chapter is devoted to supervised learning techniques for classification and regression, including nearest neighbor methods, linear and logistic regressions,
…moreIn this chapter, we present the main classic machine learning methods. A large part of the chapter is devoted to supervised learning techniques for classification and regression, including nearest neighbor methods, linear and logistic regressions, support vector machines, and tree-based algorithms. We also describe the problem of overfitting as well as strategies to overcome it. We finally provide a brief overview of unsupervised learning methods, namely, for clustering and dimensionality reduction. The chapter does not cover neural networks and deep learning as these will be presented in Chaps. 3 , 4 , 5 , and 6 .
less -
OverviewSpringer Protocols
Interpretability of Machine Learning Methods Applied to Neuroimaging
Deep learning methods have become very popular for the processing of natural images and were then successfully adapted to the neuroimaging field. As these methods are non-transparent, interpretability methods are needed to validate them and ensure
…moreDeep learning methods have become very popular for the processing of natural images and were then successfully adapted to the neuroimaging field. As these methods are non-transparent, interpretability methods are needed to validate them and ensure their reliability. Indeed, it has been shown that deep learning models may obtain high performance even when using irrelevant features, by exploiting biases in the training set. Such undesirable situations can potentially be detected by using interpretability methods. Recently, many methods have been proposed to interpret neural networks. However, this domain is not mature yet. Machine learning users face two major issues when aiming to interpret their models: which method to choose and how to assess its reliability. Here, we aim at providing answers to these questions by presenting the most common interpretability methods and metrics developed to assess their reliability, as well as their applications and benchmarks in the neuroimaging context. Note that this is not an exhaustive survey: we aimed to focus on the studies which we found to be the most representative and relevant.
less -
OverviewSpringer Protocols
Evaluating Machine Learning Models and Their Diagnostic Value
This chapter describes model validation, a crucial part of machine learning whether it is to select the best model or to assess performance of a given model. We start by detailing the main performance metrics for different tasks (classification,
…moreThis chapter describes model validation, a crucial part of machine learning whether it is to select the best model or to assess performance of a given model. We start by detailing the main performance metrics for different tasks (classification, regression), and how they may be interpreted, including in the face of class imbalance, varying prevalence, or asymmetric cost–benefit trade-offs. We then explain how to estimate these metrics in an unbiased manner using training, validation, and test sets. We describe cross-validation procedures—to use a larger part of the data for both training and testing—and the dangers of data leakage—optimism bias due to training data contaminating the test set. Finally, we discuss how to obtain confidence intervals of performance metrics, distinguishing two situations: internal validation or evaluation of learning algorithms and external validation or evaluation of resulting prediction models.
less -
OverviewSpringer Protocols
Machine Learning for Alzheimer’s Disease and Related Dementias
Dementia denotes the condition that affects people suffering from cognitive and behavioral impairments due to brain damage. Common causes of dementia include Alzheimer’s disease, vascular dementia, or frontotemporal dementia, among others. The onset
…moreDementia denotes the condition that affects people suffering from cognitive and behavioral impairments due to brain damage. Common causes of dementia include Alzheimer’s disease, vascular dementia, or frontotemporal dementia, among others. The onset of these pathologies often occurs at least a decade before any clinical symptoms are perceived. Several biomarkers have been developed to gain a better insight into disease progression, both in the prodromal and the symptomatic phases. Those markers are commonly derived from genetic information, biofluid, medical images, or clinical and cognitive assessments. Information is nowadays also captured using smart devices to further understand how patients are affected. In the last two to three decades, the research community has made a great effort to capture and share for research a large amount of data from many sources. As a result, many approaches using machine learning have been proposed in the scientific literature. Those include dedicated tools for data harmonization, extraction of biomarkers that act as disease progression proxy, classification tools, or creation of focused modeling tools that mimic and help predict disease progression. To date, however, very few methods have been translated to clinical care, and many challenges still need addressing.
less -
OverviewSpringer Protocols
A Regulatory Science Perspective on Performance Assessment of Machine Learning Algorithms in Imaging
This chapter presents a regulatory science perspective on the assessment of machine learning algorithms in diagnostic imaging applications. Most of the topics are generally applicable to many medical imaging applications, while brain disease-specific
…moreThis chapter presents a regulatory science perspective on the assessment of machine learning algorithms in diagnostic imaging applications. Most of the topics are generally applicable to many medical imaging applications, while brain disease-specific examples are provided when possible. The chapter begins with an overview of US FDA’s regulatory framework followed by assessment methodologies related to ML devices in medical imaging. Rationale, methods, and issues are discussed for the study design and data collection, the algorithm documentation, and the reference standard. Finally, study design and statistical analysis methods are overviewed for the assessment of standalone performance of ML algorithms as well as their impact on clinicians (i.e., reader studies). We believe that assessment methodologies and regulatory science play a critical role in fully realizing the great potential of ML in medical imaging, in facilitating ML device innovation, and in accelerating the translation of these technologies from bench to bedside to the benefit of patients.
less
Related Techniques
Machine Learning occurring with Expert Systems
-
ProtocolSpringer Protocols
Artificial Intelligence in Drug Safety and Metabolism
The use of artificial intelligence methods in drug safety began in the early 2000s with applications such as predicting bacterial mutagenicity and hERG inhibition. The field has been endlessly expanding ever since and the models have become more
…moreThe use of artificial intelligence methods in drug safety began in the early 2000s with applications such as predicting bacterial mutagenicity and hERG inhibition. The field has been endlessly expanding ever since and the models have become more complex. These approaches are now integrated into molecule risk assessment processes along with in vitro and in vivo methods. Today, artificial intelligence can be used in every phase of drug discovery and development, from profiling chemical libraries in early discovery, to predicting off-target effects in the mid-discovery phase, to assessing potential mutagenic impurities in development and degradants as part of life cycle management. This chapter provides an overview of artificial intelligence in drug safety and describes its application throughout the entire discovery and development process.
less -
ProtocolSpringer Protocols
Developmental Toxicity Prediction
Developmental toxicity may be estimated using commercial and noncommercial software that is already available in the market and/or literature, or models may be built from scratch using both commercial and noncommercial software packages. In this
…moreDevelopmental toxicity may be estimated using commercial and noncommercial software that is already available in the market and/or literature, or models may be built from scratch using both commercial and noncommercial software packages. In this chapter, commonly available software programs that can predict the developmental toxicity of chemicals are described. In addition, a method for developing qualitative structure–activity relationship (SAR) models to predict the developmental toxicity of chemicals qualitatively (yes/no prediction) and quantitative structure–activity relationship (QSAR) models to predict quantitative estimates (e.g., LOAEL) of developmental toxicants is also described in this chapter. Additional information described in this chapter include methods to predict physicochemical properties of chemicals that can be used as descriptor variables in the model building process, statistical methods that be used to build QSAR models as well as methods to validate the models that are developed. Most of the methods described in this chapter can be used to develop models for health endpoints other than developmental toxicity as well.
less
Machine Learning occurring with Neural Network Model
-
ProtocolSpringer Protocols
In Silico Prediction Method for Protein Asparagine Deamidation
In silico prediction methods were developed to predict protein asparagine (Asn) deamidation. The method is based on understanding deamidation mechanism on structural level with machine learning. Our structure-based method is more accurate than the
…moreIn silico prediction methods were developed to predict protein asparagine (Asn) deamidation. The method is based on understanding deamidation mechanism on structural level with machine learning. Our structure-based method is more accurate than the sequence-based method which is still widely used in protein engineering process. In addition, molecular dynamics simulation was applied to study the time occupancy of nucleophilic attack distance, which is hypothesized as the most important step toward the rate-limiting succinimide intermediate formation. A more accurate prediction method for distinguishing potentially liable amino acid residues would allow their elimination or reduction as early as possible in the drug discovery process. It is possible that such quantitative protein structure–property relationship tools can also be applied to other protein hotspot predictions.
less -
ProtocolSpringer Protocols
Application of Artificial Neural Network and Genetic Algorithm Modeling for In Vitro Regeneration of Seaweed Seedling Production
Marine macro-algae, commonly known as “seaweed,” are used in everyday commodity products worldwide for food, feed, and biostimulant for plants and animals and continue to be one of the conspicuous components of world aquaculture production. However,
…moreMarine macro-algae, commonly known as “seaweed,” are used in everyday commodity products worldwide for food, feed, and biostimulant for plants and animals and continue to be one of the conspicuous components of world aquaculture production. However, the application of ANN in seaweeds remains limited. Here, we described how to perform ANN-based machine learning modeling and GA-based optimization to enhance seedling production for implications on commercial farming. The critical steps from seaweed seedling explant preparation, selection of independent variables for laboratory culture, formulating experimental design, executing ANN Modelling, and implementing optimization algorithm are described.
less
Machine Learning occurring with KNIME
-
ProtocolSpringer Protocols
Ensemble Classifiers for Multiclass MicroRNA Classification
Gene regulation is of utmost importance to cell homeostasis; thus, any dysregulation in it often leads to disease. MicroRNAs (miRNAs) are involved in posttranscriptional gene regulation and consequently, their dysregulation has been associated with
…moreGene regulation is of utmost importance to cell homeostasis; thus, any dysregulation in it often leads to disease. MicroRNAs (miRNAs) are involved in posttranscriptional gene regulation and consequently, their dysregulation has been associated with many diseases. MiRBase version 21 contains microRNAs from about 200 species organized into about 70 clades. It has been shown that not all miRNAs collected in the database are likely to be real and, therefore, novel routes to delineate between correct and false miRNAs should be explored. We introduce a novel approach based on k-mer frequencies and machine learning that assigns an unknown/unlabeled miRNA to its most likely clade/species of origin. A simple way to filter new data would be to ensure that the novel miRNA categorizes closely to the species it is said to originate from. For that, an ensemble classifier of multiple two-class random forest classifiers was designed, where each random forest was trained on one species–clade pair. The approach was tested with different sampling methods on a dataset that was taken from miRBase version 21 and it was evaluated using a hierarchical F-measure. The approach predicted 81% to 94% of the test data correctly, depending on the sampling method. This is the first classifier that can classify miRNAs to their species of origin. This method will aid in the evaluation of miRNA database integrity and analysis of noisy miRNA samples.
less -
ProtocolSpringer Protocols
Prediction of Peptide and TCR CDR3 Loops in Formation of Class I MHC-Peptide-TCR Complexes Using Molecular Models with Solvation
Formation of major histocompatibility (MHC)-peptide-T cell receptor (TCR) complexes is central to initiation of an adaptive immune response. These complexes form through initial stabilization of the MHC fold via binding of a short peptide, and
…moreFormation of major histocompatibility (MHC)-peptide-T cell receptor (TCR) complexes is central to initiation of an adaptive immune response. These complexes form through initial stabilization of the MHC fold via binding of a short peptide, and subsequent interaction of the TCR to form a ternary complex, with contacts made predominantly through the complementarity-determining region (CDR) loops of the TCR. Stimulation of an immune response is central to cancer immunotherapy. This approach depends on identification of the appropriate combinations of MHC molecules, peptides, and TCRs to elicit an antitumor immune response. This prediction is a current challenge in computational biochemistry. In this chapter, we introduce a predictive method that involves generation of multiple peptides and TCR CDR 3 loop conformations, solvation of these conformers in the context of the MHC-peptide-TCR ternary complex, extraction of parameters from the generated complexes, and use of an AI model to evaluate the potential for the assembled ternary complex to support an immune response.
less
Machine Learning occurring with Decision Tree
-
ProtocolSpringer Protocols
Systems Biology Approaches to Understanding COVID-19 Spread in the Population
In essence, the COVID-19 pandemic can be regarded as a systems biology problem, with the entire world as the system, and the human population as the element transitioning from one state to another with certain transition rates. While capturing all
…moreIn essence, the COVID-19 pandemic can be regarded as a systems biology problem, with the entire world as the system, and the human population as the element transitioning from one state to another with certain transition rates. While capturing all the relevant features of such a complex system is hardly possible, compartmental epidemiological models can be used as an appropriate simplification to model the system’s dynamics and infer its important characteristics, such as basic and effective reproductive numbers of the virus. These measures can later be used as response variables in feature selection methods to uncover the main factors contributing to disease transmissibility. We here demonstrate that a combination of dynamic modeling and machine learning approaches can represent a powerful tool in understanding the spread, not only of COVID-19, but of any infectious disease of epidemiological proportions.
less -
ProtocolSpringer Protocols
Machine Learning to Predict Teratogenicity: Theory and Practice
Latifa DoualiMachine learning (ML) is a subfield of artificial intelligence (AI) that consists of developing algorithms that can automatically learn patterns and relationships from data, without being explicitly programmed. It continues to advance with the
…moreMachine learning (ML) is a subfield of artificial intelligence (AI) that consists of developing algorithms that can automatically learn patterns and relationships from data, without being explicitly programmed. It continues to advance with the development of more sophisticated algorithms, increased computational power, and larger datasets, leading to significant advancements in AI technology. With the significant progress made in ML, the need to apply these systems in the area of teratogenicity is growing. It is sought as robust boosting methods to overcome many limitations and restrictions facing the experimental studies. By performing tasks such as classification, regression, clustering, anomaly detection, and decision systems, ML can be used to assess whether an agent is teratogen or not or to determine its teratogenic potential. It may also be used for the purpose of deciding on the use of medicinal products. In this chapter, we describe how ML can be used to investigate teratogenicity.
less
Machine Learning occurring with Natural Language Processing
-
ArticleNature Methods
Large-scale foundation model on single-cell transcriptomics
Large pretrained models have become foundation models leading to breakthroughs in natural language processing and related fields. Developing foundation models for deciphering the ‘languages’ of cells and facilitating biomedical research is promising
…moreLarge pretrained models have become foundation models leading to breakthroughs in natural language processing and related fields. Developing foundation models for deciphering the ‘languages’ of cells and facilitating biomedical research is promising yet challenging. Here we developed a large pretrained model scFoundation, also named ‘xTrimoscFoundationα’, with 100 million parameters covering about 20,000 genes, pretrained on over 50 million human single-cell transcriptomic profiles. scFoundation is a large-scale model in terms of the size of trainable parameters, dimensionality of genes and volume of training data. Its asymmetric transformer-like architecture and pretraining task design empower effectively capturing complex context relations among genes in a variety of cell types and states. Experiments showed its merit as a foundation model that achieved state-of-the-art performances in a diverse array of single-cell analysis tasks such as gene expression enhancement, tissue drug response prediction, single-cell drug response classification, single-cell perturbation prediction, cell type annotation and gene module inference.
less -
OverviewSpringer Protocols
Knowledge Graphs and Their Applications in Drug Discovery
Knowledge graphs represent information in the form of entities and relationships between those entities. Such a representation has multiple potential applications in drug discovery, including democratizing access to biomedical data, contextualizing
…moreKnowledge graphs represent information in the form of entities and relationships between those entities. Such a representation has multiple potential applications in drug discovery, including democratizing access to biomedical data, contextualizing or visualizing that data, and generating novel insights through the application of machine learning approaches. Knowledge graphs put data into context and therefore offer the opportunity to generate explainable predictions, which is a key topic in contemporary artificial intelligence. In this chapter, we outline some of the factors that need to be considered when constructing biomedical knowledge graphs, examine recent advances in mining such systems to gain insights for drug discovery, and identify potential future areas for further development.
less
Advertisement
Broader concepts
- Machine Learning