Advertisement
Research ArticleHepatologyVirology
Open Access |  10.1172/JCI171533
10.1172/JCI171533
Hepatitis B virus infection disrupts homologous recombination in hepatocellular carcinoma by stabilizing resection inhibitor ADRM1
Ming Zeng,1 Zizhi Tang,1 Laifeng Ren,2 Haibin Wang,3 Xiaojun Wang,1 Wenyuan Zhu,4 Xiaobing Mao,5 Zeyang Li,4 Xianming Mo,5 Jun Chen,6 Junhong Han,5 Daochun Kong,4 Jianguo Ji,4 Antony M. Carr,7 and Cong Liu1
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Zeng, M. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Tang, Z. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Ren, L. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Wang, H. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Wang, X. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Zhu, W. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by
Mao, X.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Li, Z. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by
Mo, X.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by
Chen, J.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by
Han, J.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Kong, D. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Ji, J. in: JCI | PubMed | Google Scholar
1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by
Carr, A.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pediatrics, Key Laboratory of Birth Defects and Related Diseases of Women and Children (Ministry of Education), West China Second University Hospital, Sichuan University, Chengdu, China.
2Department of Immunology, Shanxi Hospital Affiliated to Cancer Hospital, Chinese Academy of Medical Sciences, Taiyuan, China.
3Department of Pediatric Surgery, Wuhan Children’s Hospital, Huazhong University of Science and Technology, Wuhan, China.
4State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Beijing, China.
5State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu, China.
6MOE Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, College of Life Sciences, Zhejiang University, Hangzhou, China.
7Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Brighton, United Kingdom.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Find articles by Liu, C. in: JCI | PubMed | Google Scholar
Authorship note: MZ, ZT, LR, and HW contributed equally to this work.
Published October 10, 2023 - More info
J Clin Invest. 2023;133(23):e171533. https://doi.org/10.1172/JCI171533.
© 2023 Zeng et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: April 18, 2023; Accepted: October 5, 2023
-
Abstract
Many cancers harbor homologous recombination defects (HRDs). A HRD is a therapeutic target that is being successfully utilized in treatment of breast/ovarian cancer via synthetic lethality. However, canonical HRD caused by BRCAness mutations do not prevail in liver cancer. Here we report a subtype of HRD caused by the perturbation of a proteasome variant (CDW19S) in hepatitis B virus–bearing (HBV-bearing) cells. This amalgamate protein complex contained the 19S proteasome decorated with CRL4WDR70 ubiquitin ligase, and assembled at broken chromatin in a PSMD4Rpn10- and ATM-MDC1-RNF8–dependent manner. CDW19S promoted DNA end processing via segregated modules that promote nuclease activities of MRE11 and EXO1. Contrarily, a proteasomal component, ADRM1Rpn13, inhibited resection and was removed by CRL4WDR70-catalyzed ubiquitination upon commitment of extensive resection. HBx interfered with ADRM1Rpn13 degradation, leading to the imposition of ADRM1Rpn13-dependent resection barrier and consequent viral HRD subtype distinguishable from that caused by BRCA1 defect. Finally, we demonstrated that viral HRD in HBV-associated hepatocellular carcinoma can be exploited to restrict tumor progression. Our work clarifies the underlying mechanism of a virus-induced HRD subtype.
-
Introduction
Long-term liver infection with hepatitis B virus (HBV) predisposes carriers to hepatocellular carcinoma (1). Therapy for late-stage HBV-associated hepatocellular carcinoma remains a major challenge, with poor therapy responses and low overall survival (2). HBV subverts the Cullin4-DDB1-RING (CRL4) ubiquitin ligase for viral purposes via the oncoprotein HBx (3). Among the CRL4 subcomplexes that subsequently show reduced function is CRL4WDR70, a regulator for homologous recombination and chromatin remodeling (4). In HBV+ cells, enrichment of homologous recombination (HR) factors including RPA, RAD51, and EXO1 at DNA double-strand breaks (DSBs) is compromised, correlated to reduced CRL4WDR70 activity and DNA end resection (5). However, the mechanistic role of CRL4WDR70 in DNA repair remains undetermined.
Misregulation of DSB repair compromises chromosomal stability (6) and is often characterized by altered usage of non-homologous end joining (NHEJ) and HR (7). DNA end processing by 5′-to-3′ resection governs HR commitment by generating RPA-coated single-stranded DNA (ssDNA) that subsequently loads RAD51 to form a filament that enables homology search (8). The cell cycle position and the chromatin context surrounding the DSB site influence ssDNA production (9, 10). DSBs occurring in G1 are repaired by NHEJ, while those occurring after replication are repaired by NHEJ and/or HR. The choice of pathway is regulated by the competitive occupancy of 53BP1 and BRCA1 at DNA breaks (11–13). A new therapeutic strategy, synthetic lethality (SL), has recently been introduced for cancer subtype–specific chemotherapy, and this was first exemplified by the treatment of homologous recombination defect (HRD) breast cancers carrying BRCA1/2 mutations (BRCAness) with PARP inhibitors (PARPis), including olaparib and talazoparib (14).
DNA repair is also regulated by the 19S regulatory particle (RP), a constituent of the 26S proteasome that degrades ubiquitin-tagged proteins. Distinct from the protease activity sequestered in 20S core particle (CP), the canonical 19S RP recognizes ubiquitinated targets and deubiquitinates and positions them for translocation and unfolding to allow degradation by the CP (15). The 19S RP is subdivided into a “base” that is constituted of 6 paralogous AAA+ ATPases (PSMC1–6) plus several non-ATPase proteins (PSMD1Rpn2, PSMD2Rpn1, and ADRM1Rpn13) and a “lid” containing PCI domain proteins (PSMD3, 6, 8, 11–13), MPN domain proteins (PSMD7Rpn8 and POH1Rpn11), plus the non-PCI/MPN domain subunits including DSS1 (also known as Rpn15 or Sem1) (16, 17). The base and lid are conformationally dynamic and together bind a further subunit, the ubiquitin receptor PSMD4Rpn10.
As introduced above, in addition to regulating proteolysis, the RP also performs non-proteolytic roles in the context of chromatin. This was originally identified from the recruitment of a subset of RP proteins to the GAL1-10 promoter, implying a direct role in RNA polymerase II transcription (18). In the context of DNA repair, the 19S has subsequently been shown to modulate the efficiency of both DSB and nucleotide excision repair (19–21). DSS1Sem1 and POH1Rpn11 locate at DSB sites and regulate repair activities of RAD52/Pol4 and 53BP1/RAP80, respectively (22–25). The number and diversity of 19S-associating proteins and functions have obscured the elucidation of its mechanism in chromatin biology, and a comprehensive model depicting its interplay with DNA repair machinery is lacking.
Here we provide evidence that CRL4WDR70 forms a specific complex with the break-associated 19S proteasome (subsequently referred to as CDW19S; CULLIN4A-DDB1-WDR70-19S) that favors HR via end processing. 19S RP controls both MRE11 and EXO1 nucleases, and CRL4WDR70 engages with an EXO1-specific module of RPs to catalyze the ubiquitination and degradation of ADMR1Rpn13, a 19S-associated ubiquitin receptor that we identify as a resection barrier factor. We show that HBx disintegrates CUL4A-DDB1 from CDW19S and leaves scaffold-free WDR70-19S on damaged DNA. As such, HBx retards the clearance of ADRM1Rpn13 from DNA breaks, whereby a special HRD subtype is produced distinct from that caused by BRCA1 deficiency. Similar to the addiction of BRCAness cells to PARP functions, the viral HRD sensitizes HBV+ cells to PARPi. These data uncover a viral HRD subtype resulting from failed clearance of resection inhibitors.
-
Results
HBx induces HRD by perturbing the balanced choice of DSB repair. To assess the impacts of HBV/HBx on DSB repair, an I-SceI–induced DSB system was used to measure sister chromatid repair (HR), single-strand annealing (SSA), and NHEJ efficiencies (26). T43 hepatocytes, which harbor integrated HBV genomes (5), displayed a modest decrease in HR and SSA relative to the parental HBV-free L02 cells (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/JCI171533DS1). To establish whether this HBV-induced HR deficiency can be attributed, at least in part, to HBx ablating CRL4WDR70, we expressed HBx in HEK293T cells and in WDR70-knockout derivatives (Figure 1A). HBx biased repair away from HR/SSA and toward NHEJ as expected. WDR70 deletion did not affect the repair profile beyond that seen when HBx is expressed in 293T cells, consistent with an epistatic relationship.
 Figure 1
Figure 1Interference with CRL4WDR70 by HBx induces a viral HRD. (A) Repair frequency of indicated pathways in WDR70-knockout or HBx-expressing cells relative to control cells (293T). **P <0.05 by 2-tailed t test. NS, no statistic significance. (B) Left: Example confocal images showing 53BP1 (red) and RPA32 (green) IRIF in HBx-expressing L02 cells 8 hours after IR. Soluble nuclear proteins were preextracted with 0.1% Triton X-100. Scale bar: 10 μm. Right: Pixel intensity (vertical) across the maximal central line of individual IRIFs. Precipitation of red line (53BP1) and rising of green line (RPA32) along the vertical axis indicate the central cavity. (C and D) ChIP assay depicting p-RPA32 chromatin loading at indicated distance from the DSB upon expression of gRNA (g1) targeting the PPP1R12C/p84 locus. WDR70-knockout or control 293T cells (C) and L02 cells expressing HBx (D) were cotransfected with si53BP1 or control siRNA (siScr). (E) Relative HR/SSA efficiency for L02 cells pretreated with HBx, siWDR70, or siBRCA1 and concomitant silencing of 53BP1. (F and G) Representative images (F) and quantification (G) of aberrant chromosomes in the indicated cells cotreated with olaparib (1 μM) and/or sh53BP1. (H) Giemsa staining for colony formation (left) and survival curves (right) of control L02 (HBV–) and T43 (HBV+) cells subjected to olaparib treatment. Survival at endpoints was analyzed for statistical significance. n = 3 biological repeats; error bars indicate SD; t test. (I) Survival curves for L02 and T43 cells treated with siWDR70 (left) or sh53BP1 (right) with simultaneous exposure to indicated concentrations of olaparib. n = 3 experimental repeats; error bars indicate SD; P values by t test are shown for indicated groups.
53BP1 establishes a resection barrier at DSB ends after ionizing radiation (IR). To activate HR, 53BP1 is subsequently displaced from the focal center of ionizing radiation–induced foci (IRIF) in a BRCA1-dependent manner (23). This 53BP1 loss correlates with the accumulation of ssDNA binding factor (RPA32) within the central cavity of the focus, a process that is dependent on functional resection. Upon HBV infection, HBx expression, or WDR70 loss, a reduction in the size of central cavities of 53BP1 IRIF that is positive for RPA32 was observed, indicative of impaired displacement of 53BP1 by pro-HR factors (Figure 1B and Supplemental Figure 1B). Consistent with the antagonistic role of BRCA1 against 53BP1 (11), HBx expression or direct compromising of CRL4WDR70 diminished attraction of BRCA1 to DSBs (Supplemental Figure 1C).
HR defects caused by BRCA1 mutations can be rescued by removal of the 53BP1-mediated resection barrier (27). Similar to this situation, in HBx-expressing cells, depleting 53BP1 restored camptothecin-induced (CPT-induced) RPA32 phosphorylation (p-RPA32) respective to controls (Supplemental Figure 1D), and similarly restored IRIF of p-RPA32 and RAD51 recombinase in cells transfected with small interfering RNA of WDR70 (siWDR70) (Supplemental Figure 1E). We evaluated DSB loading of 53BP1 in WDR70-knockout and HBx-expressing cells using chromatin immunoprecipitation (ChIP) assay developed for CRISPR-induced DSBs at the PPP1R12C/p84 locus (5), where enhanced 53BP1 binding was detected at 0.5–10 kb from the break in comparison with their respective controls (Supplemental Figure 1F). When 53BP1 was concomitantly knocked down in WDR70-knockout and HBx-expressing cells, restricted p-RPA32 association within 6–10 kb from the break site was reactivated to levels comparable to those in wild-type cells (Figure 1, C and D). Resembling the revitalization of BRCAness HRD by 53BP1 depletion, simultaneous silencing of 53BP1 restored HR/SSA efficacies in HBx expression and siWDR70 cells (Figure 1E). We conclude that HBV/HBx-induced viral HRD shares the common defect with BRCAness HRD in terms of 53BP1 accumulation.
In the context of BRCA1 ablation, preferential channeling of DSB repair to error-prone NHEJ promotes the generation of toxic chromosomal structures that are exacerbated upon PARP inhibition (27, 28). Similarly, chromosomal aberrations were synergistically exacerbated in HBx-expressing L02 and T43 cells upon PARPi (olaparib) addition, and this was suppressible by 53BP1 ablation (Figure 1, F and G). Strikingly, T43 cells were hypersensitive to a range of PARPis when compared with HBV-free L02 cells (Figure 1H and Supplemental Figure 1G), revealing a strong synthetic lethality between HBV/HBx and PARP inhibition. A second HBV+ cell line (HepG2.2.15) displayed comparable SL phenotype relative to its parental line (HepG2; Supplemental Figure 1H). This SL subtype was linked to CRL4WDR70, as siWDR70 sensitized L02 cells to PARPi but showed no additional effect on T43 cell PARPi sensitivity (Figure 1I, left). T43 sensitivity to PARPi was rescued by sh53BP1 (Figure 1I, right), reinforcing the contribution of DSB-associated 53BP1 in viral HRD induction.
Assembling CDW19S on break-associated chromatin. To establish how viral HRD is induced and CRL4WDR70 interplays with the repair machinery, we purified proteins associated with TAP-tagged Wdr70 from fission yeast, given the evolutionary conservation of CRL4WDR70 (4). Ddb1, the adaptor that links Wdr70 and Pcu4 (Cullin4 homolog), was deleted to disable the ubiquitin ligase activity, thus preventing the degradation of binding factors. MALDI-TOF mass spectrometry of sliced gel bands identified 3 categories of proteins (Figure 2A and Supplemental Table 1): histone species (Htb1H2B, Hht1H3.1), ubiquitination factors (note we saw low coverage for Pcu4 and abundant ubiquitin), and, unexpectedly, a range of proteins derived from 19S RP of the proteasome. The panel of RP subunits encompasses the main RP subcomplexes including PCI, MPN, and ATPase domain proteins. Notably, no peptides from the proteolytic 20S core particle were retrieved.
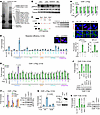 Figure 2
Figure 2CDW19S engages DSB-proximal chromatin. (A) TAP-affinity-purified spWdr70-interacting proteins separated by gradient SDS-PAGE. Proteins identified by MALDI-TOF mass spectrometry are shown on the right. See peptide coverage in Supplemental Table 1. Subdomains of interface (Int), PCI, MPN, and ATPase (ATP) are indicated for RP subunits using human and yeast nomenclatures. IgH, heavy chain of rabbit IgG; TEV, tobacco etch virus endopeptidase. (B) Immunoblotting for p-RPA32 and H2B monoubiquitination (uH2B) in L02 cells with indicated siRNA and CPT treatment. (C) Left: Illustration for XbaI-based resection assay at the selected AsiSI-dependent DSB site (chromosome 1: 89,458,595–89,458,603). Right: Example of monitoring of ssDNA production by semiquantitative PCR. (D) Quantification for XbaI-based resection assay showing DSB processing in DIvA cells depleted for the indicated CDW19S subunits. Inset: Excessive p-RPA32 immunostaining implies hypersection in siADRM1Rpn13 RPE1 cells. Data normalized to control (siScr) with 4-OHT induction. Original magnification (inset): ×400. (E) ChIP assay 2.5 kb distal to an AsiSI-induced DSB showing break association of FLAG-tagged CDW19S subunits upon PSMD4Rpn10 ablation relative to control transfection. DIvA cells all treated with 4-OHT. (F) Left: ChIP assay for FLAG-tagged WDR70/DDB1 at indicated distances from AsiSI-induced DSB ends. Right: Representative PCR products. (G) ChIP assay of FLAG-PSMD4Rpn10 2.5 kb from an AsiSI-induced DSB upon PSMD4WT or PSMD4dUIM expression. Anti-FLAG immunoblotting is shown in the right panel. (H) Equivalent ChIP assay for indicated CDW19S subunits in the presence of PSMD4WT or PSMD4dUIM expression. (I) Top: Representative images of p-RPA32 and BRCA1 IRIF in the presence of PSMD4Rpn10 or PSMD4dUIM (4 hours after IR). Nuclei counterstained with DAPI. Scale bars: 10 μm. Bottom: Quantification of fluorescent intensity or foci numbers. In H and I, PSMD4 plasmids are FLAG-less and siRNA resistant, and cells were cotransfected with siPSMD4Rpn10. (J and K) PSMD4Rpn10 enrichment upon 4-OHT induction at 0.5 kb from an AsiSI-induced DSB after treatment with the indicated siRNA or inhibitors.
Coimmunoprecipitation for WDR70/DDB1 in cells ectopically expressing FLAG-tagged 19S subunits solidified the conservation of CRL4WDR70-19S interaction in human cells (Supplemental Figure 2A). Consistent with a role in DSB repair, these interactions were enhanced upon CPT treatment (Supplemental Figure 2B). A functional importance of the interaction is implied by the observation that, as shown previously for HBx expression and siWDR70 (5), depleting the proteasomal PSMD2Rpn1 subunit inhibited pro-HR events including RPA32 phosphorylation and H2B monoubiquitylation (Figure 2B). We thus conclude that CRL4WDR70 and the 19S particle form a complex (CDW19S) that influences DSB repair.
To survey the function of the CDW19S complex at DSBs, the DIvA system, where DNA breaks are generated by ER-tagged AsiSI endonuclease upon its nuclear import following 4-OHT treatment (29), was used to analyze a specific AsiSI site on chromosome 1 (89,458,595–89,458,603, reference genome hg19) by ChIP. Four hours after 4-OHT treatment, p-RPA32 was observed in control cells between 0.5 and 5 kb from the break, whereas loss of WDR70 affected distal (2.5–5 kb) but not proximal (0.5–1 kb) deposition (Supplemental Figure 2C). Distal resection (3.3 kb) was further analyzed by digestion of genomic DNA with XbaI, which is inactive on ssDNA (Figure 2C). Uncut ssDNA was quantified by PCR across the digestion site, and 4 hours after induction, control cells showed evidence of ssDNA at 1 kb and 3.3 kb from the break site. In this resection system, knockdown of the majority of CDW19S subunit genes, plus those encoding the associated chaperones (PSMD5Hsm3 and PSMD9Nas2) and PSMD10 proteins, exhibited siWDR70-like resection defects relative to siScramble (Figure 2D). ADRM1Rpn13 depletion stood out, displaying hyperactive ssDNA production relative to control cells. No function for PSMD12Rpn5, PSMC5Rpt6, and PSMC6Rpt4 was observed, despite significant mRNA ablation (Supplemental Figure 2D). Moreover, ChIP assay revealed that all RP and WDR70/DDB1 proteins (FLAG-tagged) were enriched at 2.5 kb distal to DSBs following AsiSI nuclear import (Figure 2, E and F). We conclude that CRL4WDR70 decorates 19S RP in a stable CDW19S complex to stimulate resection.
PSMD4Rpn10 recruits CDW19S in an ATM-MDC1-RNF8–dependent manner. 19S RP is known to be targeted to chromatin and interplay with DSB-associated ubiquitin conjugates (24, 30). We speculated that the ubiquitin binding directs the RP to the DSBs. PSMD4Rpn10, an integral ubiquitin receptor in the RP lid that recruits K63- or K48-linked ubiquitin targets to the proteasome, is a promising candidate (31). Knockdown of PSMD4Rpn10 abolished DSB recruitment of WDR70 and all RP subunits except ADRM1Rpn13 (Figure 2E). In contrast, PSMD4Rpn10 was recruited to DSBs irrespective of siRNA treatment for any of the CDW19S subunits tested (Supplemental Figure 2E). PSMD4Rpn10 recognizes ubiquitin chains via a ubiquitin-interacting motif (UIM), mutation of which abrogates ubiquitin affinity (32). Ectopically expressed UIM-deletion mutant (PSMD4dUIM) dissociated from DSBs (Figure 2G) and suppressed DSB assembly of CDW19S complex (Figure 2H). Consistent with this, PSMD4dUIM expression phenocopied siWDR70, repressing resection as monitored by p-RPA32 and BRCA1 IRIF (Figure 2I).
To establish whether damage signaling was prerequisite for CDW19S DSB association, we evaluated the abundance of PSMD4Rpn10 at break sites when early damage signaling was interfered with. Silencing the damage-responsive E3 enzyme (RNF8) and its partner E2 (UBC13), rather than RNF168 or RNF20, effectively abolished PSMD4Rpn10 loading 0.5 kb from the DSB (Figure 2J). RNF8-UBC13 catalyzes K63-polyubiqutination chains, suggesting that PSMD4Rpn10 is attracted to DSB-associated K63-modified proteins. The RNF8 FHA domain docks to TQxF clusters on MDC1 following their ATM-dependent phosphorylation (33). Consistent with this, both ATM kinase activity (kinase inhibitor KU55933) and MDC1 (siMDC1) were required for the DSB attraction of PSMD4Rpn10 (Figure 2K). We conclude that the assembly of CDW19S is initiated at break proximity by PSMD4Rpn10 recognition of K63 species that is ATM-MDC1-RNF8 dependent.
CDW19S is functionally segregated into MRE11- and EXO1-regulatory modules. Long-range resection is mainly promoted by exonuclease 1 (EXO1) and suppressible by 53BP1. To establish whether CDW19S has a functional domain mediating extensive resection, individual subunits were knocked down, and p-RPA32 recruitment was assayed 0.5 and 2.5 kb from an AsiSI site following 4-OHT induction. As seen with siWDR70, ablation of the majority of 19S components led to p-RPA32 reduction at 2.5 kb. For many components, this was effectively reverted by concomitant si53BP1 treatment (Figure 3A). The effects of siPSMD4Rpn10 and siPSMD5Hsm3 on RPA32 loading were reproduced with a second siRNA, and each of these was complemented by respective siRNA-resistant plasmids (Supplemental Figure 3A). Interestingly, the distal resection defects observed upon PSMD2Rpn1, PSMD4Rpn10, and MPN (POH1Rpn11 and PSMD7Rpn8) depletion were not restored by si53BP1. This correlates to inability of 53BP1 knockdown to rescue p-RPA32 reduction at 0.5 kb of these mutants (Figure 3B and Supplemental Figure 3B), documenting the role of these 4 components in initiating proximal resection that does not require CRLWDR70 or RP subunits with WDR70-like phenotypes.
 Figure 3
Figure 3Separate CDW19S modules regulate MRE11 and EXO1 activation. (A and B) ChIP assay showing DSB loading of p-RPA32 at 2.5 kb (A) or 0.5 kb (B) distal to an AsiSI-induced DSB upon silencing of indicated CDW19S subunits. Concomitant 53BP1 knockdown was performed in A. (C) Enumeration of MRE11 foci upon silencing of individual CDW19S subunits in RPE1 cells. Immunofluorescence was carried out 30 minutes after IR. n = 3 biological repeats, 50 cells counted for each repeat. Error bars indicate SD. P values by t test are shown. (D) ChIP assay showing loading of EXO1 at 2.5 kb distal to an AsiSI-induced DSB.
To further explore these distinctions, we analyzed the formation of MRE11 IRIF. CRL4WDR70 depletion or HBx expression does not diminish the proximal nuclease (MRE11) from DNA ends (5). However, the ablation of PSMD2Rpn1, PSMD4Rpn10, PSMD7Rpn8, or POH1Rpn11 prevented the formation of MRE11 IRIF 30 minutes and 2 hours after IR (Figure 3C and Supplemental Figure 3, C and D). This reflects their requirement for both proximal (0.5 kb) and distal (2.5 kb) processing (Figure 3, A and B). We therefore categorize an MRE11-regulatory module (MRM) within CDW19S that is necessary to initiate end processing.
We next analyzed chromatin association of the long-range exonuclease EXO1. Distal EXO1 association (2.5 kb) was reduced upon silencing of WDR70 and CDW19S subunits that phenocopy WDR70 (Figure 3D). The same impact was not observed upon silencing ADRM1Rpn13 and non-functional components including PSMD12Rpn5, PSMC5Rpt6, or PSMC6Rpt4. Again, that siADRM1Rpn13 promotes distal EXO1 and p-RPA32 recruitment relative to control cells suggests that ADRM1Rpn13 limits, rather than promotes, long-range resection (Figure 2D and Figure 3, A and D). We conclude that CRL4WDR70, together with most RP components, constitutes an EXO1-specific module (ESM) that regulates extensive resection.
PSMD1Rpn2, ADRM1Rpn13, PSMD5Hsm3, and PSMD9Nas2 are required for CRL4WDR70 recruitment. To establish how CRL4WDR70 docks to the RP, WDR70 ChIP was exploited in the DIvA system combined with targeted siRNA. CRL4WDR70 loading at 2.5 kb following AsiSI nuclear import was significantly compromised by PSMD1Rpn2, ADRM1Rpn13, PSMD5Hsm3, and PSMD9Nas2 ablation (Figure 4A), implying that a platform dependent on these 4 ESM subunits tethers CRL4WDR70 to the RP. Ablation of other subunits of MRM (PSMD2Rpn1, POH1Rpn11) and ESM (PSMD3Rpn3, PSMC1Rpt2, POH1Rpn11) did not exhibit dramatic impact. Further, coimmunoprecipitation between WDR70 and FLAG-PSMD4Rpn10 in chromatin fractions was abolished in the absence of PSMD1Rpn2, ADRM1Rpn13, and PSMD9Nas2, but was maintained upon depletion of representative interface (PSMD2Rpn1), PCI (PSMD3Rpn3), MPN (POH1Rpn11), or ATPase (PSMC1Rpt2) subunits (Figure 4B).
 Figure 4
Figure 4The docking platform of CRL4WDR70 on 19S RP. (A) ChIP assay for FLAG-tagged WDR70 at 2.5 kb distal to an AsiSI-induced DSB following siRNA treatment of 19S subunits. Data normalized to siScramble with 4-OHT induction. (B) Coimmunoprecipitation of FLAG-PSMD4Rpn10 and WDR70 from chromatin fractions of CPT-treated HEK293T cells with or without ablation of indicated RP components. (C) Schematic showing the high-salt procedure for screening 19S components mediating direct engagement with WDR70. (D) Pull-down assay using purified WDR70 and PSMD5Hsm3. (E and F) In vitro pull-down assay for purified WDR70 (0.5 μg) and 19S proteasome (2 μg), the latter containing FLAG-UCHL5. Recombinant PSMD5Hsm3 (1 μg, E) or specific antibodies (0.5 μg, F) were added into the reaction. (G and H) ChIP assay for loading of FLAG-tagged PSMD5Hsm3 (G) or DDB1/WDR70 (H) 2.5 kb distal to an AsiSI-induced DSB following siRNA treatments. (I) Immunoprecipitation for endogenous WDR70 and FLAG-tagged 19S subunits with or without PSMD5Hsm3 silencing.
To identify a direct docking site of CRL4WDR70, 19S particles were dissociated in nuclear HEK293T extracts using high-salt buffer (600 mM NaCl) and incubated with His-WDR70 (112–654 aa) purified from Sf9 cells (Figure 4C). The sole putative WDR70 binding partner identified was PSMD5Hsm3, a chaperone that contributes to RP assembly and only loosely associates with mature 19S (34). This interaction was confirmed by coprecipitation of recombinant His-WDR70 and Strep-PSMD5Hsm3 (Figure 4D and Supplemental Figure 4A). Recombinant WDR70 also displayed affinity with purified 19S RP (R&D Systems, E-367) containing FLAG-UCHL5 (Figure 4E and Supplemental Figure 4B). This WDR70-19S interaction is attributable to a low amount of PSMD5Hsm3 that copurified with 19S (Supplemental Figure 4C), since it was boosted by the addition of additional (0.5 μg) recombinant PSMD5Hsm3 and was weakened by pretreatment with specific antibodies against either PSMD5Hsm3 or WDR70 (Figure 4, E and F). As with other CDW19S subunits, PSMD5Hsm3 deposition at DSB sites required PSMD4Rpn10 but not WDR70 or DDB1 (Figure 4G), and siPSMD5Hsm3 impeded the DSB association of both DDB1 and WDR70 (Figure 4H). siPSMD5Hsm3 also attenuated coimmunoprecipitation of endogenous WDR70 with FLAG-PSMD4Rpn10, PSMD12Rpn5, ADRM1Rpn13, and POH1Rpn11 (Figure 4I). PSMC6Rpt4 was mildly affected. Thus, PSMD5Hsm3, together with ADRM1Rpn13, PSMD1Rpn2, and PSMD9Nas2, chaperones CRL4WDR70 to the RP, reminiscent of the chaperone contribution to the stepwise assembly of RP base subcomplexes (34).
Hereby, we posit that PSMD4Rpn10 launches the DSB assembly of CDW19S, triggering proximal and distal resection via separate modular functions of MRM and ESM by regulating nuclease activities of MRE11 and EXO1. Commitment to long-range resection requires the engagement of CRL4WDR70 to ESM via PSMD5Hsm3 (Supplemental Figure 4D).
CRL4WDR70 regulates the ubiquitin-dependent degradation of break-associated ADRM1Rpn13. CRL4WDR70 is a ubiquitin ligase that promotes resection, suggesting that it targets an anti-resection factor for degradation and implicating that such a factor would be more abundant on broken chromatin in the absence of CDW19S. ADRM1Rpn13 is excessively associated with chromatin in the absence of CDW19S (see Figure 2E), and its ablation results in increased resection (see Figure 2D) and RPA/EXO1 loading (see Figure 3, A and D). We therefore tested ADRM1Rpn13 recruitment to an AsiSI-induced DSB by ChIP, with or without siRNA of WDR70. ADRM1Rpn13 was recruited more abundantly upon siWDR70, particularly at the distal (2.5 kb) region (Supplemental Figure 5A). Importantly, co-depletion of ADRM1Rpn13 and WDR70 restored EXO1 recruitment at 2.5 kb when compared with WDR70 depletion alone (Supplemental Figure 5B).
Like PSMD4Rpn10, ADRM1Rpn13 is an RP ubiquitin receptor (35), and its DSB recruitment occurs independently of CDW19S (see Figure 2E). However, ADRM1Rpn13 supports the loading of WDR70 as part of the recruiting platform (see Figure 4A), implying their proximity. ADRM1Rpn13 encodes a C-terminal DEUBAD domain (265–407 aa) and a conserved N-terminal pleckstrin-like receptor for ubiquitin (Pru) (1–150 aa; ref. 35) that preferably binds to K48 polyubiquitin chains via a triple-residue motif. Substitution of I75R, F76R, and D79N (ADRM1mIFD) abolished the ubiquitin affinity (Supplemental Figure 5C), and impaired ADRM1Rpn13 enrichment at AsiSI-dependent DSBs (Supplemental Figure 5D). Two different ADRM1Rpn13 siRNAs restored resection in the absence of WDR70, as measured by p-RPA32 ChIP (Figure 5A, left). Coexpression of siRNA-resistant wild-type ADRM1Rpn13 abolished the rescue, whereas the mIFD version did not (Figure 5A, right), suggesting that ubiquitin association via the Pru domain is required for the anti-resection function. Therefore, CRL4WDR70 counteracts ADRM1Rpn13, and loss of ADRM1Rpn13 function obviates the need for CRL4WDR70 to promote extensive resection.
 Figure 5
Figure 5CRL4WDR70 targets ADRM1Rpn13 for UPS degradation. (A) Left: ChIP assay showing DSB loading of p-RPA32 at 2.5 kb distal to DSB upon silencing of ADRM1Rpn13 (2 different siRNAs) or WDR70. Right: Equivalent assay expressing si001-resistant wild-type (WT) or mIFD mutant ADRM1Rpn13. (B) Protein abundance of ADRM1Rpn13 in chromatin fraction of 293T cells as measured by immunoblotting (see also Supplemental Figure 5E). WDR70 was ablated, followed by CPT insult (2 μM) for 1 hour and release into drug-free medium. (C) Immunoblotting for ADRM1Rpn13 in cells cotreated with CPT and MG132. (D–F) Ubiquitin pull-down assay to identify polyubiquitinated species of FLAG-ADRM1Rpn13 upon treatment with siWDR70 (D), expression of ubiquitin variants (E), or ADRM1Rpn13 K>R mutants (F). Cells were challenged with 2 μM CPT for 2 hours. (G) Immunoblotting for CPT-induced p-RPA32 upon expression of WT, K99R, or K99-only mutant of FLAG-ADRM1Rpn13 that is siADRM1Rpn13 resistant. (H) Reconstitution of FLAG-ADRM1Rpn13 ubiquitination catalyzed by purified proteins. (I) Equivalent reconstitution using WT or K99R versions. GF-ADRM1, GST/FLAG-tagged ADRM1. (J and K) ADRM1Rpn13 ubiquitination reconstituted with addition of purified 19S and/or His-PSMD5HSM3 (J), or in the presence of anti-WDR70 or anti-PSMD5HSM3 (K). (L) ChIP assay for DSB-associated ADRM1Rpn13 (2.5 kb from an AsiSI-induced DSB; top panel) and immunoblotting for chromatin-bound ADRM1Rpn13 following CPT treatment with or without PSMD5HSM3 silencing (bottom panel). (M) Schematic for chromatin regulation of DSB repair by coordinative action of CDW19S modules.
To further test whether ADRM1Rpn13 was being targeted for degradation via CRL4WDR70, stability of chromatin-associated ADRM1Rpn13 was evaluated. ADRM1Rpn13 was depleted from the chromatin fraction in CPT-challenged 293T cells, but remained stable when cotreated with siWDR70 (Figure 5B and Supplemental Figure 5E) or proteasomal inhibitor (MG132; Figure 5C), revealing a WDR70-dependent and ubiquitin-proteasome system–dependent (UPS-dependent) turnover. Furthermore, polyubiquitination of chromatin-associated ADRM1Rpn13 was decreased upon siWDR70 (Figure 5D) and impaired by the expression of a K48R, but not a K63R, ubiquitin mutant (Figure 5E).
Seven lysine residues conserved between human and yeast ADRM1Rpn13 (Schizosaccharomyces pombe has 2 homologs, Rpn13a and Rpn13b) were identified as putative ubiquitination targets (Supplemental Figure 5F). Surveying ubiquitination profiles in cells coexpressing individual FLAG-tagged K>R mutations and HA-tagged ubiquitin identified that mutation of K99 (ADRM1K99R), but not other lysine residues (i.e., K97), abrogated ubiquitin-conjugated ADRM1Rpn13 species in the chromatin fraction (Figure 5F). Strikingly, an ADRM1K99-only mutant was sufficient for ADRM1Rpn13 polyubiquitination, and this remained WDR70 dependent (Supplemental Figure 5G). Consistent with K99 determining ADRM1Rpn13 ubiquitination, CPT-induced p-RPA32 level was reduced when siADRM1Rpn13 was complemented with ADRM1K99R relative to that seen with either wild-type or ADRM1K99-only (Figure 5G). Collectively, these results support the hypothesis that CRL4WDR70 promotes the ubiquitination and degradation of chromatin-bound ADRM1Rpn13.
19S complex boosts ADRM1Rpn13 ubiquitination via CRL4WDR70-PSMD5HSM3 engagement. To corroborate the direct ADRM1Rpn13 ubiquitination by CRL4WDR70, the CRL4WDR70 E3 ligase was reconstituted using purified subunits (His-tagged CULLIN4A, DDB1, ROC1, and truncated WDR70112–654) coexpressed in Sf9 insect cells (Supplemental Figure 5H). The artificial CRL4WDR70 complex was supplemented with a ubiquitin reconstitution system (activating enzyme [E1], conjugation factor [E2, UbcH5b], and biotinylated ubiquitin) and tested for in vitro ubiquitinylation of bacterially expressed FLAG-ADRM1Rpn13 (Supplemental Figure 5I). Ubiquitin conjugates were observed with slow-migrating mono- and polyubiquitination species (Figure 5H), absent in control reaction conducted with ADRM1K99R (Figure 5I). Consistent with the in vivo modification, ADRM1Rpn13 conjugates were K48-linked and were catalyzed by UbcH5b, but not UbcH5a or UbcH6 (Supplemental Figure 5, J and K).
In vivo, the loading platform containing ADRM1Rpn13 and PSMD5Hsm3 potentially juxtaposes the CRL4WDR70 E3 ligase to substrates (i.e., ADRM1Rpn13). We thus examined how purified 19S affected CRL4WDR70-dependent ADRM1Rpn13 ubiquitination. Ubiquitination activity was substantially stimulated by supplementing of RP particles to the CRL4WDR70-ADRM1Rpn13 (GST/FLAG-tagged) reconstitution system. This was further boosted by concomitant addition of recombinant His-PSMD5Hsm3 (Figure 5J) and inhibited by inclusion of anti-PSMD5 or anti-WDR70 antibodies (Figure 5K). Consistent with this, in vivo siPSMD5Hsm3 resulted in accumulation of ADRM1Rpn13 at an AsiSI-induced DSB and stabilization after CPT treatment (Figure 5L). Taking together the results from Figures 2–5, we propose that ADRM1Rpn13 pauses end resection at DSB proximity, a function that relies on PSMD4Rpn10 and MRM. Additionally, we propose that to activate EXO1 and extensive resection, CRL4WDR70 acts as an RP-associated E3 ligase to catalyze UPS-mediated removal of ADRM1Rpn13, a function sustained by the PSMD5Hsm3-containing subdomain within ESM (Figure 5M).
HBx attenuates the ESM to generate a viral HRD. HBx disintegrates the CRL4WDR70 complex by displacing WDR70 from CUL4A-DDB1 via a biomimetic H-box motif (5, 36). In HBV-integrated T43 hepatocytes PSMD2Rpn1 maintained its association with WDR70 while partially dissociating from DDB1, in contrast to its interaction with both proteins in HBV– L02 cells (Supplemental Figure 6A). Given that DDB1 association with damaged chromatin is WDR70 dependent (4), we reasoned that HBx detaches the CRL4-DDB1 scaffold from ESM, leaving partially assembled CDW19S (WDR70-19S) at breaks. Indeed, expressing HBx in DIvA cells pronouncedly reduced DDB1 loading at AsiSI-induced DSBs without affecting other CDWS19 components (Figure 6A).
 Figure 6
Figure 6Torso CDW19S and ADRM1Rpn13 accumulation marks HBV-induced HRD subtype. (A) ChIP assay (left) for FLAG-tagged CDW19S subunits 2.5 kb from DSB in the presence or absence of HA-tagged HBx expression (right). Quantification was normalized to DDB1 value without HBx expression. (B) Chromatin and soluble nuclear fractionation of indicated proteins upon CPT insult with or without HBx-HA expression. Densitometry for DDB1 from 3 repeats is shown on the right. Results were obtained from identical biological samples immunoblotted from different concentrations of PAGE gels. (C) Enrichment of EXO1 or p-RPA32 loading 2.5 kb distal from DSB with HBx expression or siADRM1Rpn13. (D) HR/SSA repair assay in the presence of HBx or siWDR70 in L02 cells, with or without concomitant siADRM1Rpn13 treatment. (E) Schematic showing the HBx-induced “torso” CDW19S and consequent failure of ADRM1Rpn13 removal. (F) Representative images (left) and counting (right) of 53BP1 IRIF (8 hours after IR) in WDR70-ablated cells. Simultaneous ADRM1Rpn13 silencing was performed as indicated. Scale bar: 10 μm. (G) ChIP showing the inability of siADRM1Rpn13 to restore the DSB loading of p-RPA32 and EXO1 in BRCA1-depleted cells. (H) Parallel comparison of HR/SSA improvement by control siRNA (green) or siADRM1Rpn13 (orange) in BRCA1- and WDR70-depleted cells. P values for multiple-group comparison in B–D, F, and G were calculated by 2-way ANOVA test.
As a result, HBx impedes the EXO1-specific function of CDW19S. HBx expression in L02 cells accumulated chromatin-bound ADRM1Rpn13 but decreased DDB1 association upon CPT treatment (Figure 6B), an observation reproducible in another HBV+ HepG2.2.15 cell line relative to its parental line (HepG2; Supplemental Figure 6B). HBx reduced attachment of p-RPA32 and EXO1 distal (2.5 kb) to an AsiSI-induced DSB (Figure 6C), and this was not exacerbated when PSMD4Rpn10 (ubiquitin-dependent DSB anchorage) or PSMD5Hsm3 (ESM subunit) was concomitantly depleted (Supplemental Figure 6C). As with WDR70 deficiency, siADRM1Rpn13 reversed the HBx-induced impediment of p-RPA32/EXO1 loading. As we reported previously, HBx does not affect MRE11 kinetics (5). Thus, HBx leaves “torso” CDW19S complex lacking the CUL4A-DDB1 scaffold at DSBs, and affects extensive resection by disrupting CRL4WDR70-containing ESM.
The impact of ADRM1Rpn13 accumulation and torso CDW19S was further evaluated by analysis of the repair outcomes in I-SceI–induced DSB system. Imbalanced activities of NHEJ/HR in the presence of HBV or HBx were epistatic with siRNA of either PSMD4Rpn10 or PSMD5Hsm3 (Supplemental Figure 6, D and E). As expected, the HBx-dependent defect in pathway choice (HR and particularly SSA) was, in part, reversed by siADRM1Rpn13 (Figure 6D). Again, ADRM1Rpn13 inhibition corrected repair bias in HepG2.2.15 cells (Supplemental Figure 6F). Collectively, torso CDW19S resulting from HBx induces HR defects by maintaining DSB-bound ADRM1Rpn13 and impeding EXO1-dependent resection (Figure 6E).
ADRM1Rpn13 accumulation distinguishes viral HRD from BRCAness HRD. We continued to establish the functional link between ADRM1Rpn13 and 53BP1 regarding their roles in resection barrier. Foci analysis revealed that ADRM1Rpn13 depletion reduced 53BP1 IRIF in WDR70-defective cells (Figure 6F), whereas depletion of 53BP1 exerted no effect on DSB-associated ADRM1Rpn13 in siWDR70 or HBx-expressing cells (Supplemental Figure 6G). Apparently, ADRM1Rpn13 acts upstream to erase the 53BP1-mediated resection barrier. Given the rescuing effects of si53BP1 on HRD in the context of torso CDW19S, these results indicate that 53BP1 absence can bypass the demand of ADRM1Rpn13 degradation for the commitment of long-range resection. Unlike the ability of 53BP1 depletion to rescue BRCAness HRD (see Figure 1E), siADRM1Rpn13 failed to restore the distal loading of p-RPA32 and EXO1, and to reverse the HR/SSA defects in BRCA1-depleted cells (Figure 6, G and H). Thus, while viral HRD and BRCA1-defective HRD share the common defect of 53BP1 accumulation, only the viral subtype is driven by excessive ADRM1Rpn13 deposition at DSBs.
Targeting HBV-induced HRD suppresses the disease progression of HBV-associated hepatocellular carcinoma. The overall survival of HBV-associated hepatocellular carcinoma (HBVHCC) patients is low, and few contemporary chemotherapeutic treatments are widely applicable. We exploited the viral HRD–induced SL and tested the potential of PARPi treatment in tumor-burdened immunodeficient mice. In athymic nude mice implanted with T43 xenografts, tumor growth was strongly restricted by 131.5 mg/kg/d olaparib monotreatment (Figure 7A and Supplemental Figure 7A). Notably, low doses of olaparib and cisplatin imposed synergistic effects on both cellular viability and T43 xenografts (Figure 7, B and C). To establish the benefits of PARPi treatment for HBVHCC, patient-derived xenografts were implanted in immunocompromised NOD-Prkdcscid-IL2rg(em1-IDOM) mice that subsequently were subjected to treatment with clinically relevant and mouse-equivalent dosage of olaparib (100 mg/kg/d) (37), in conjunction with a low dose of cisplatin (0.5 mg/kg/2 days). Cisplatin-insensitive hepatocellular carcinoma (HCC) xenografts from 4 HBV+ patients (patients 16, 17, 19, and 23) and 1 HBV-free patient (patient 76) were subjected to 1 course of olaparib/cisplatin (O/C) conjunctive treatment. Body weights in both O/C and vehicle groups were monitored throughout the experiment (Supplemental Figure 7B). Compared with the unrestricted tumor growth in the vehicle-only group, all HBVHCC cases treated with O/C displayed tumor growth inhibition (from 95.17% to 51.09%) at terminal therapy (Figure 7D), reflecting significantly delayed tumor progression (Figure 7E and Supplemental Figure 7C). Furthermore, O/C treatment produced a significantly longer median period of progression-free survival compared with that of vehicle groups (P = 0.003; Figure 7F). In contrast, xenografts derived from HBV-free HCC (patient 76) exhibited no therapy response when subjected to the same course of O/C treatment (Figure 7D).
 Figure 7
Figure 7HBV-induced HRD subtype sensitizes HBVHCC to PARP inhibition. (A) Responses of T43 xenografts to monotreatment with olaparib or vehicle. Error bars indicate SD; t test. (B and C) Responses of T43 cells (B, 3 biological repeats) and xenografts (C, 6 littermates included) to conjunctive administration of olaparib and cisplatin. Tumor volumes are presented as means ± SD. DMSO: equivalent amount of solvent solution. Error bars indicate SD; P values were calculated by t test (B) and by 2-way ANOVA test (C). (D) Schematic of PARPi administration to HCC engraftment in NOD-SCID mice (top) and tumor responses (bottom). Tumor volumes of 4 HBVHCCs (patients 16, 17, 19, and 23) and 1 HBV-free HCC (patient 76, progressive disease) are shown at indicated days after inoculation. Olaparib: 33.3 mg/kg/d; cisplatin: 0.5 mg/kg/2 days (O/C). Horizontal axis, days after tumor transplantation; arrows, starting date of medication. Numbers of animals were as follows: patient 16 (vehicle, n = 4; treatment, n = 4), patient 17 (vehicle, n = 2; treatment, n = 2), patient 19 (vehicle, n = 3; treatment, n = 2), patient 23 (vehicle, n = 3; treatment, n = 2), patient 76 (vehicle, n = 3; treatment, n = 3). (E) Tumor response for HBVHCC patient-derived xenograft (PDX) sublines treated with vehicle or O/C at week 2–3. Graphs show mean ± SEM, analyzed with 2-sided unpaired Student’s t test. (F) Kaplan-Meier plot indicating progression-free survival of HBVHCC sublines. The y axis is the percentage of animals whose tumor volumes were smaller than 300 mm3. P value was calculated by log-rank test.
-
Discussion
In addition to regulating proteolysis, the 19S regulatory particle has a variety of non-proteolytic functions. Among these are a range of activities in the context of chromatin, including transcription initiation and elongation (18, 38, 39), roles in histone modification (40), and DNA repair (19–25). In this study we identify an RP decorated with CRL4WDR70 (CDW19S) that assembles in the vicinity of DSBs. We show that CDW19S can be segregated into distinct functional modules defined by ubiquitin anchorage (PSMD4Rpn10) and subdomains required for the resection activity of MRE11 (MRM) and EXO1 (ESM) nucleases. Their differential functions at proximal (0.5 kb) and distal (2.5 kb) indicate a central role of 19S RP in coordinating proximal and distal DNA end processing.
Mechanistically, we identified ADRM1Rpn13 as a direct catalytic substrate for CRL4WDR70 associated with the RP and show that preventing ADRM1Rpn13 degradation by mutating a single lysine residue (K99>R) attenuates distal resection, underpinning ADRM1Rpn13 as a key inhibitor of long-range resection upstream of 53BP1. Intriguingly, the equivalent K99 residue only exists in one of the fission yeast ADRM1 homologs, Rpn13b, and it was this protein that copurified with WDR70 (Figure 2A and Supplemental Figure 5F). This suggests a functional divergence between fission yeast Rpn13 paralogs in the context of UPS and 19S RP chromatin functions in low eukaryotes, while in human cells both functions are carried out by a single ADRM1Rpn13 version. We propose that CDW19S, upon association with RNF8-catalyzed K63 species, engages its associated enzymatic activities (i.e., CRL4WDR70) to remove the ADRM1Rpn13-dependent barrier and promotes long-range resection via EXO1 activation (Figure 5M).
Our uncovering of a pathological accumulation of ADRM1Rpn13 in the presence of HBV/HBx raises the intriguing question of how ADRM1Rpn13 functions in physiological contexts. In eukaryotes, resection is a vital process for HR and thus genome stability. However, end processing should also be tightly controlled to avoid overproduction of ssDNA structures (41) highly susceptible to nuclease attack (i.e., APOBEC/AID), as well as resulting mutagenesis (42, 43). From this angle, the pro- and counter-HR actions of CRL4WDR70-ADRM1Rpn13 constitute a quality control for appropriate DNA processing. Alternatively, as described in a recent study, 53BP1 may foster fidelity of homology-dependent repair by suppressing the switch from error-free gene conversion by RAD51 to mutagenic SSA by RAD52 (44). How ADRM1Rpn13 operates in these scenarios alongside the BRCA1-53BP1 control of pathway choice remains an interesting question.
The HBV oncoprotein HBx plays versatile roles during HBVHCC development, including redirecting the Cullin4-DDB1 scaffold to degrade SMC5/6 complex that counteracts viral replication (3). By hijacking the DDB1-containing ubiquitin ligases, HBx causes a deficit in the cellular ligases that rely on the Cullin4-DDB1 scaffold, including CRL4WDR70 and its associated function of HR (5). Here we clarify the underlying mechanism by which HBV/HBx pathogenesis causes HRD, showing that HBx prevents the integration of Cullin4-DDB1 into chromatin-associated CDW19S. This torso CDW19S is unable to promote ADRM1Rpn13 degradation, thus compromising homology-dependent repair. Notably, although this viral HRD subtype affects BRCA1-53BP1 function, it is different from canonical BRCAness HRD that does not involve the malfunction of ADRM1Rpn13.
BRCAness HRD renders cells sensitive to PARP inhibition as has been applied in breast cancer treatment (45, 46). In vitro studies suggest that HCC cell lines are also susceptible to PARP inhibition (47), corresponding to elevated levels of PARP1/2 enzymes and PARylation in HCC tissues (48, 49). While these studies did not indicate a clear HRD phenotype that deminishes the viability of HBVHCC cells through PARPi, we showed that HBV/HBx-induced viral HRD is SL to PARP inhibition, which relied on the impairment of CDW19S. We further demonstrate, using xenograft models of T43 cells or patient-derived HBVHCC materials, that PARPi treatment effectively constrains tumor growth and improves survival. This underlines the potential for SL treatment of HBVHCC using HRD toxins.
For breast cancer, HRD rates are estimated by either BRCAness mutations or single-nucleotide variation (signature 3) via whole-genome sequencing (WGS) (50). It would be worthwhile to determine the incidence and genomic features of viral HRD by WGS, which would not only provide the biomarkers for PARPi treatment of HBVHCC, but also help discriminate the fingerprints between BRCAness and HBV HRD subtypes.
Taken together, our results identify the mechanism by which HBV causes a special HRD subtype, and establish that HBVHCC is an HRD cancer type susceptible to PARP inhibition. Our findings provide a mechanistic justification for targeting HBVHCC by SL.
-
Methods
Cell culture and reagents. Human cell lines (HEK293T, WDR70KO, RPE1, DIvA, HepG2 and HepG2.2.15, L02, and T43) were maintained in culture media supplemented with 10% FBS. L02 and T43 cells were propagated from single clones. The STR profile and hepatic identity of L02 are shown in Supplemental Figure 8, and T43 cells with stably integrated HBV genomes were regularly selected in G418 (200 μg/mL) and examined for titration of HBs and HBe antigens by ELISA (5). RPE1 and DIvA were obtained from the cell bank of the Genome Damage Stability Centre, University of Sussex. L02, HEK293T, HepG2, and HepG2.2.15 were purchased from National Infrastructure of Cell Line Resource (Wuhan or Shanghai, China). All cell lines tested negative for mycoplasma contamination. Plasmid and siRNA transfections were carried out using Lipofectamine 3000 (Invitrogen, L3000015). DSBs in DIvA system were induced by 300 nM 4-OHT (MilliporeSigma, H7904) for 4 hours. For ATM kinase or proteasome inhibition, 10 μM KU55933 (Selleck, S1092) or 100 μM MG132 (Selleck, S2619) was used for pretreatment for 2 hours. Plasmids, Primers, antibodies, and siRNA used in this study are listed in Supplemental Tables 2–5.
Plasmids. For cloning of lentiviral vectors for expression of FLAG-tagged proteins, PCR fragments were inserted into the EcoRI site of pLVX-G-FLAG plasmid using the In-Fusion cloning kit (Clontech, 639650). Plasmids used in this study are listed in Supplemental Table 2.
Evaluation of gene expression. For detection of gene expression, 106 cells were harvested and total RNA was extracted using NucleoZOL Reagent (Macherey-Nagel, 740404). cDNA was obtained from 1 μg RNA by Reverse Transcription System (Promega, A3500). Real-time quantitative PCR (qPCR) and semiquantitative PCR were performed on a Bio-Rad CFX96 Real Time System according to the instructions of qPCR Kits (Promega, A6001) with 3 technical replicates for each sample. For reverse transcriptase qPCR, the relative expression was calculated by fold change of target gene normalized to GAPDH/18S rRNA in each sample, and the experimental group was normalized to the average of the control. Semiquantitative PCR products were resolved by 2% agarose electrophoresis.
Tandem affinity purification for Wdr70-associating proteins. To purify proteins in complex with Wdr70 of S. pombe, 8 liters of spWdr70 tandem affinity purification–tagged (TAP-tagged) or control strains were grown in YEP rich media (1% Yeast Extract [Solarbio, Y8020], 2% Peptone [Solarbio, P8450], 0.5% NaCl). The cell pellet was washed and resuspended in an equal volume of lysis buffer (25 mM Tris-HCl [pH 7.5], 15 mM EGTA, 15 mM MgCl2, 0.1% NP-40, 1 mM DTT, 0.1 mM NaF) supplemented with protease inhibitor (Complete Mini, Roche, 11836153001) plus DTT (1 μM) and subjected to mechanical beating with glass beads in a ribolyzer. The granular cell pellets and the powder of ground cells were frozen at –80°C and stored (up to 3 months) before use. Soluble extracts were clarified by centrifugation before binding to rabbit IgG-coated Dynabeads M280 (Invitrogen, 11203D), followed by extensive washing and cleavage by tobacco etch virus (TEV) protease. Purified Wdr70-associating components were resolved by 8%–20% gradient SDS-PAGE and stained with Brilliant Blue G-Colloidal Concentrate Kit (MilliporeSigma, B-2025).
Excised gel bands were washed with 25 mM NH4HCO3 containing 50% acetonitrile and dehydrated with 100% acetonitrile, followed by treatment with 10 mM DTT and incubation for 1 hour at 65°C. After cooling, gel samples were alkylated with 55 mM iodoacetamide for 45 minutes at room temperature. Samples were digested with trypsin (1:50, wt/wt) dissolved in 25 mM NH4HCO3 at 37°C overnight. Digested peptides were extracted with 5 mM octyl-β-d-glucopyranoside in 0.25% trifluoroacetic acid for 60 minutes at 37°C and directly applied onto the AnchorChip target (Bruker Daltonics), which was loaded with α-cyano-4-hydroxycinnamic acid (CHCA) thin-layer matrix. Mass spectra of extracted peptides for each sample were determined using Ultraflex MALDI-TOF/TOF mass spectrometry (Bruker Daltonics) in a positive ion reflector mode. The ion acceleration voltage was 25 kV. Both MALDI-TOF spectra and the MS/MS spectra were processed by FlexAnalysis 2.2 (Bruker) and Biotool 2.2 (Bruker) and automatically searched against the Swiss-Prot database using Mascot software (Matrix Science). Main parameters were set as follows: mass range from 800 to 4,000 Da; S/N ≥ 3.0; fixed modification, carbamidomethyl (Cys); variable modification, oxidation (Met); maximum number of missing cleavages, 1; MS tolerance, 50 ppm; and MS/MS tolerance, 0.7 Da.
Protein purification and pull-down assay. Purified 6xHis-WDR70 (112–654 aa) and CRL4-WDR70 tetraplex from Sf9 insect cells were purchased from HitGen. For PSMD5 purification, the coding sequence was amplified from a cDNA library and cloned into the pET28a-2xStrep plasmid. For bacterial expression, BL21 strain transformed with expression vector was induced with 1 mM IPTG overnight at 16°C. Five hundred milliliters of cells were pelleted and resuspended in 20 mL TBST buffer with protease inhibitor cocktail and sonicated. Centrifuged supernatant was incubated with MagStrep XT Beads (IBA Lifesciences, 2-4090-002) and eluted with BXT buffer (100 mM Tris [pH 8.0], 150 mM NaCl, 1 mM EDTA, 50 mM biotin) for 1 hour. For ADRM1 wild-type or K99R purification, pET28a-FLAG plasmids were used with otherwise similar protocol except that incubation was with GSH beads (BeaverBeads, 70601-5) and elution with buffer B (50 mM Tris-HCl, 10 mM glutathione).
For immunoprecipitation, 106 cells were harvested and lysed in 200 μL of buffer A (10 mM HEPES [pH 7.9], 10 mM KCl, 1.5 mM MgCl2, 0.34 M sucrose, 10% glycerol, 1 mM DTT, 0.1% Triton X-100, and protease inhibitors). Nuclei were collected in the pellet by low-speed centrifugation (1,500g, 4 minutes, 4°C) and further incubated with buffer B (2 mM EDTA, 0.2 mM EGTA, 1 mM DTT, and protease inhibitor mixture) for 10 minutes on ice. After centrifugation (2,000g, 4 minutes), the pellet was resuspended in IP2 buffer (50 mM Tris [pH 8.0], 150 mM NaCl, 1% NP-40, and protease inhibitor cocktail) with 100 U Micrococcal Nuclease (New England Biolabs, M0247S)/DNase I (New England Biolabs, M0303S) and incubated on ice for 30 minutes to digest genome DNA. The chromatin fraction was clarified by high-speed centrifugation (21,000g, 10 minutes) and then incubation with prewashed FLAG M2 beads (MilliporeSigma, M8823) to pull down FLAG-tagged immunocomplex. Proteins were detected by immunoblotting.
Immunostaining. Indirect immunofluorescent staining was described elsewhere (5). Briefly, cryosectioned tissues or cells on coverslips were fixed with 4% PFA. Permeabilized sections were incubated with primary antibodies and labeled with secondary antibodies (anti-rabbit–Cy3 or anti-mouse–FITC), followed by mounting with anti-fade medium containing DAPI (Vector Laboratories, H120010) and visualization using a Leica fluorescence microscope (DM4 M) or Olympus fluorescence microscope (BX51). All quantitative immunostaining analysis was performed by counting of 100–200 cells from 3 independent experiments.
For measurement of 53BP1 exclusion from the core IRIF, images were processed using FV10-ASW 3.1 Viewer (Olympus) software, and the cavity of 53BP1 and sizes of p-RPA32 foci were gauged by measurement of pixel densities by ImagePro Plus 6.0 (Media Cybernetics) across the center lines of foci with best visualization. Fifty to two hundred foci were measured for each group.
In vitro ubiquitination assay. Recombinant E2 enzymes (R&D Systems, K-980B) and ubiquitin conjugation initiation kit (R&D Systems, K-995) were used for ubiquitination assay. In brief, 500 ng purified ADRM1 or mutant protein was incubated with UBE1, UbcH5b, ubiquitin, CRL4WDR70 tetramer (500 ng) at 37°C for 1 hour. The samples were mixed with SDS sample buffer and detected by immunoblotting following SDS-PAGE. Recombinant ubiquitin mutants K48R (R&D Systems, UM-K48R) and K63R (R&D Systems, UM-K63R) were used for identification of ubiquitin chain species. Recombinant 19S proteasome with FLAG-tagged UCHL5 was purchased from R&D Systems (E-367).
Measurement for efficacies of DSB repair. For plasmid-based DSB repair system (26), pCMV plasmids containing the I-SceI restriction site were subject to thorough in vitro digestion. Cut plasmids (NHEJ and SSA: 5 μg; HR: 10 μg; gift from Jun Chen, Zhejiang University, Hangzhou, China) were transfected into 293T or WDR70-knockout cells and allowed to repair in vivo for 48 hours. Recombined or ligated plasmids by SSA/HR (primers 3/4) or NHEJ (primers 5/4) were recovered from cells by a HighPure PCR Template Preparation Kit (Roche, 11796828001), followed by quantitative real-time PCR with appropriate primers (Supplemental Table 3) specific for SSA/HR or NHEJ fragments using a Bio-Rad CFX96 Real Time System. The amount of extracted plasmid DNA was normalized to the product of primers 1/2. The relative frequency of each repair pathway was defined as fold changes relative to qPCR values of repaired DNA extracted in parallel from wild-type or mock-infected 293T cells.
For the XbaI-dependent resection assay, genomic DNA from 2 × 105 DIvA cells following 4-OHT induction was purified using High Pure PCR Template Preparation (Roche, 11796828001). For each sample, 300 ng of extracted DNA was digested with XbaI at 37°C for 4 hours, and the reaction was stopped by heating at 65°C for 10 minutes. For ssDNA quantification, 20 ng digested sample was amplified by qPCR using primers listed in Supplemental Table 3.
Chromatin immunoprecipitation. Chromatin immunoprecipitation (ChIP) assays were performed as previously described with minor modifications (51). Briefly, for each location reaction, approximately 4 × 106 cells were harvested and then cross-linked for 10 minutes with formaldehyde at a final concentration of 1% and washed twice in PBS. Cells were then lysed in lysis buffer (50 mM HEPES [pH 7.4], 140 mM NaCl, 1% Triton X-100, 0.1% Na deoxycholate, 1 mM EDTA supplemented with protease inhibitor cocktail [Roche, 11836170001]) and sonicated to solubilize chromatin and shear the cross-linked DNA. Sonication was performed at 4°C with Bioruptor Pico (Diagenode) at default power for 40 30-second long pulses (30-second pause between pulses). To retrieve chromatin-associated proteins, the whole-cell extracts were incubated on a rotator overnight at 4°C with 10 μL magnetic Protein G Dynabeads (Invitrogen, 10004D) precoated with 2 μg of the indicated antibodies or FLAG M2 beads (MilliporeSigma, M8823). The beads were washed sequentially for 5 min with lysis buffer (see above), lysis high-salt buffer (50 mM HEPES [pH 7.4], 500 mM NaCl, 1% Triton X-100, 0.1% Na deoxycholate, 1 mM EDTA), wash buffer (10 mM Tris [pH 8.0], 250 mM LiCl, 0.5% NP-40, 0.5% Na deoxycholate, 1 mM EDTA), and TE buffer (pH 8.0), and twice for each buffer. Bound DNA-immune complexes were eluted off the beads in elution buffer (50 mM Tris [pH 8.0], 1% SDS, 10 mM EDTA) by heating at 65°C for 2 hours. DNA was purified using HighPure PCR Template Preparation Kit (Roche, 11796828001). Whole-cell extracts were treated in parallel for cross-link reversal and DNA extraction. Two hundred nanograms recovered DNA was used for each PCR reaction using Platinum Taq DNA Polymerase (Invitrogen, 10966). All reactions were performed in triplicate. Primer sets used to measure protein enrichment were as follows: CRISPR system (input, pair 22/23; 0.5 kb, pair 6/7; 3.5 kb, pair 8/9; 10 kb, pair 18/19); DIvA system (0.5 kb, pair 26/27; 1 kb, pair 28/29; 2.5 kb, pair 30/31; 5 kb, pair 32/33).
Preparation of metaphase chromosomal spread. Cells were plated in a 60-mm dish and arrested in mitosis by 2-hour treatment with colcemid (final concentration of 200 ng/mL). Prewarmed 75 mM KCl was then added to trypsinized cells and incubated for 15 minutes at 37°C. Then 4 drops of freshly prepared fixative (3:1 solution of methanol/acetic acid) were added. Cells were pelleted and resuspended in 5 mL fixative and incubated for 20 minutes at 4°C. After the fixing step was repeated 3 or 4 times, cell pellets were resuspended in 0.5 mL fixative solution. Two or three drops of fluid were precipitated onto a prechilled slide from a height of 18 inches. Slides were air-dried thoroughly and stained using the Giemsa protocol. The mitotic chromosomes were observed and evaluated using an Olympus fluorescence microscope (BX51) at ×1,000 magnification.
Cell survival assay. For colony formation assays, cells were plated at 500 cells each in a 10-cm dish and cultured at 37°C for 10–14 days, then fixed with methanol and stained with Giemsa solution. PARPi (olaparib/KU0059436, talazoparib/S7048, Selleck; niraparib/MB5556, veliparib/MB5524, rucaparib/MB1643, Meilunbio) or nutlin (ApexBio, A4228) was applied in the indicated concentrations alone or simultaneously used with diluted cisplatin (Supertrack Bio-pharmaceutical, 131102). Cell proliferation of WDR70-knockdown 293T cells was measured by cell counting every other day after transfection.
PARP inhibition using nude mice and xenograft model. All animal experiments were carried out in animal facilities of West China Second University Hospital or at an outsourcing service (Beijing IDMO Co. Ltd.). Athymic nude immunodeficient mice (BALB/c nu/nu, specific pathogen free) were purchased from the animal center of Sichuan University. For T43 xenograft mouse models, female nude mice starting at 4–5 weeks of age were used for the experiments. To assess the tumorigenicity of L02 or T43 cells in vivo, mice were subcutaneously inoculated in both sides of armpits or hind flanks (4 × 106 cells per site). When T43 xenografts had reached an average volume of approximately 0.01 cm3, animals were randomized (using a random number table) into treatment groups, and 5 animals were selected for each group. Monotreatment with olaparib was administered intraperitoneally to each animal at a dosage of 131.5 mg per kg body weight per day. For double treatment, olaparib was administered at a dosage of 41 mg per kg body weight per day, and cisplatin at 0.42 mg per kg body weight every 2 days. A parallel group of mice was given PBS-diluted DMSO as control. Inoculated and drug-administered mice were observed each day. Sizes of tumors were measured by Vernier caliper and tumor volumes calculated by 3D measurement (length × width × height × π/6) until termination of the experiments. To exclude outliers of tumor volume, Grubbs’s test was applied (Gi>GO. 95, 2-tailed test). Experiments were performed blindly.
Patient-derived xenograft assays were established and performed in specific pathogen–free facilities of Beijing IDMO Co. Ltd. Primary HBVHCC tissues were enzymatically dissociated, and primary tumor cells were subcutaneously inoculated into the flank of NOD-Prkdcscid-IL2rg(em1-IDOM) (NPI) recipient mice. After sufficient tumor growth, tumors were passaged or cryopreserved for banking. For PARPi treatment of HBVHCC, 4 cisplatin-insensitive HBVHCCs and 1 HBV-free HCC were inoculated subcutaneously to NSG mice at 4–6 weeks of age. Randomized and age-matched males were used with an even split between control and drug administration. In total, 22 tumor-burdened mice survived to the terminal stage of the experiments, and data points were collected from these animals. Each tumor included a vehicle and an experimental group, to which 2–3 repeats were allocated, performed blindly. Vehicle (12.5% DMSO in PBS) or combined olaparib (33.3 mg/kg/d) plus cisplatin (3 mg/kg/2 days) treatment was administered by intraperitoneal injection. The health of the animal was monitored daily throughout therapy. Sizes of xenograft implantations were regularly measured by Vernier caliper and calculated for 3D volumes. Mean percentages of inhibition were calculated by % inhibition = (mean(control) – mean(treatment))/mean(control) × 100%. HBV+ and HBV– HCC materials used in patient-derived xenograft experiments were selected according to the HBV infection status: HBV+ tumors were serologically HBsAb–HBsAg+HBcAb+HBeAb+HBeAg–, and HBV– tumors were HBsAb+HBsAg–HBcAb–HBeAb–HBeAg–. All HCCs were from Chinese male patients.
Data analysis. The percentage of tumor growth inhibition was defined as [1 – (mean volume of treated tumors)/(mean volume of control tumors)] × 100%. Progression-free survival (PFS) was defined in this study as the condition where tumor volumes were under 300 mm3. Mice with tumor sizes exceeding 300 mm3 were assigned “progressive disease” and taken off PFS calculation. These mice could be enrolled again into PFS calculation if tumor regressed to less than 300 mm3 during later treatment. The PFS curves were generated by the Kaplan-Meier method, and log-rank tests were used to compare PFS in each experimental group.
Statistics. All histograms are presented as means ± SD. For quantitative analysis including ChIP assay, image analysis, and repair analysis, at least 3 independent experiments were carried out. Kaplan-Meier plots and log-rank tests were computed by SPSS 16.0 software to compare the survival outcomes between 3 groups. P values were calculated by 2-tailed Student’s t test between 2 groups, or by 2-way ANOVA test for multiple-group comparison, using GraphPad Prism 6. P values less than 0.05 were considered significant. Endpoint values of cell survival and antitumor assays were used for statistical analysis.
Study approval. Animal studies were approved by ethical committees of West China Second University Hospital, Chengdu, China (Medical Research 2018(015)) and Beijing IDMO Co. Ltd., Beijing, China (P20211229001), and were performed strictly in compliance with the ethical guidelines and regulations. Animals were housed in accordance with approved protocols, and efforts were made to minimize suffering. Deidentified tumor material was collected from patients in agreement with institutional ethical regulations. Informed consent was provided via protocol [Medical Research 2018(014)] approved by the ethical committees of West China Second University Hospital.
Data availability. Data included in this article are provided in the Supporting Data Values file and are available upon request from the corresponding authors. Participant data are available upon request from CL (congliu@scu.edu.cn) under standard rules of data protection and ethical permissions.
-
Author contributions
CL purified CDW19S complex, and MZ characterized it. LR and HW analyzed 53BP1 and BRCA1 in the absence of CRL4WDR70. JC assisted with I-SceI repair assay. XW performed immunoblotting and cell culture. WZ, ZL, and JJ identified components by mass spectrometry. X Mo advised on animal models. ZT investigated PARPi toxicity and anticancer studies. X Mao and JH contributed CRL4WDR70-mediated ADRM1 degradation. DK, AMC, and CL designed and supervised the research and interpreted data. CL and AMC acquired funding and edited and reviewed the manuscript. Co–first authors were assigned based on the academic contribution of individual authors: characterization of CDW19S architecture and mechanism by MZ, exploration of targeted therapy by ZT and the establishment of CRL4WDR70 in 53BP1 removal (LR and HW).
-
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31771580, 82073027, 81702795, 81972592, and 81821002), Sichuan University (2020SCUNL204 and SCU2019C4198), the Department of Science and Technology of Sichuan Province (2019JDTD0020, 2020YFH0019, and 2022NSFSC1502), and the 1.3.5 project for disciplines of excellence of West China Hospital (ZYGD20007, ZYJC18011, ZYGD18003, and ZYJC21021). AMC acknowledges Medical Research Council award G1100074.
Address correspondence to: Cong Liu, West China Second University Hospital, Sichuan University, Ren Ming Nan Lu, No. 17, Chengdu 610041, China. Phone: 0086.28.85501715; Email: congliu@scu.edu.cn. Or to: Antony M. Carr, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, BN1 9RQ, United Kingdom. Phone: 0044.1273.678122; Email: carr.am@sussex.ac.uk. Or to: Jianguo Ji, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.10.62755470; Email: jijg@pku.edu.cn. Or to: Daochun Kong, State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences, Peking University, Yiheyuan Lu No. 5, Haidian District, Beijing 100871, China. Phone: 0086.62760866; Email: kongdc@pku.edu.cn. Or to: Junhong Han, State Key Laboratory of Biotherapy, West China Hospital, Ren Ming Nan Lu, No. 17, Sichuan University, Chengdu 610041, China. Phone: 0086.028.61528677; Email: hjunhong@scu.edu.cn.
-
Footnotes
MZ’s present address is: Centre for Translational Research in Cancer, Sichuan Cancer Hospital and Institute, Chengdu, China.
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2023, Zeng et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: J Clin Invest. 2023;133(23):e171533. https://doi.org/10.1172/JCI171533.
-
References
- Llovet JM, et al. Hepatocellular carcinoma. Nat Rev Dis Primers. 2021;7(1):6.
- Levrero M, Zucman-Rossi J. Mechanisms of HBV-induced hepatocellular carcinoma. J Hepatol. 2016;64(1 suppl):S84–S101.
- Decorsière A, et al. Hepatitis B virus X protein identifies the Smc5/6 complex as a host restriction facto74r. Nature. 2016;531(7594):386–389.
- Zeng M, et al. CRL4(Wdr70) regulates H2B monoubiquitination and facilitates Exo1-dependent resection. Nat Commun. 2016;7:11364.
- Ren L, et al. The antiresection activity of the X protein encoded by hepatitis virus B. Hepatology. 2019;69(6):2546–2561.
- Hoeijmakers JH. DNA damage, aging, and cancer. N Engl J Med. 2009;361(15):1475–1485.
- Cao L, et al. A selective requirement for 53BP1 in the biological response to genomic instability induced by Brca1 deficiency. Mol Cell. 2009;35(4):534–541.
- Zhao F, et al. DNA end resection and its role in DNA replication and DSB repair choice in mammalian cells. Exp Mol Med. 2020;52(10):1705–1714.
- Escribano-Díaz C, et al. A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol Cell. 2013;49(5):872–883.
- Mirman Z, et al. 53BP1-RIF1-shieldin counteracts DSB resection through CST- and Polα-dependent fill-in. Nature. 2018;560(7716):112–116.
- Becker JR, et al. BARD1 reads H2A lysine 15 ubiquitination to direct homologous recombination. Nature. 2021;596(7872):433–437.
- Hu Q, et al. Mechanisms of BRCA1-BARD1 nucleosome recognition and ubiquitylation. Nature. 2021;596(7872):438–443.
- Densham RM, Morris JR. Moving mountains-the BRCA1 promotion of DNA resection. Front Mol Biosci. 2019;6:79.
- Slade D. PARP and PARG inhibitors in cancer treatment. Genes Dev. 2020;34(5–6):360–394.
- Budenholzer L, et al. Proteasome structure and assembly. J Mol Biol. 2017;429(22):3500–3524.
- Unverdorben P, et al. Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome. Proc Natl Acad Sci U S A. 2014;111(15):5544–5549.
- Bai M, et al. In-depth analysis of the lid subunits assembly mechanism in mammals. Biomolecules. 2019;9(6):213.
- Gonzalez F, et al. Recruitment of a 19S proteasome subcomplex to an activated promoter. Science. 2002;296(5567):548–550.
- Jacquemont C, Taniguchi T. Proteasome function is required for DNA damage response and Fanconi anemia pathway activation. Cancer Res. 2007;67(15):7395–7405.
- Gillette TG, et al. The 19S complex of the proteasome regulates nucleotide excision repair in yeast. Genes Dev. 2001;15(12):1528–1539.
- Russell SJ, et al. The 19S regulatory complex of the proteasome functions independently of proteolysis in nucleotide excision repair. Mol Cell. 1999;3(6):687–695.
- Krogan NJ, et al. Proteasome involvement in the repair of DNA double-strand breaks. Mol Cell. 2004;16(6):1027–1034.
- Kakarougkas A, et al. Co-operation of BRCA1 and POH1 relieves the barriers posed by 53BP1 and RAP80 to resection. Nucleic Acids Res. 2013;41(22):10298–10311.
- Butler LR, et al. The proteasomal de-ubiquitinating enzyme POH1 promotes the double-strand DNA break response. EMBO J. 2012;31(19):3918–3934.
- Tsolou A, et al. The 19S proteasome subunit Rpn7 stabilizes DNA damage foci upon genotoxic insult. IUBMB Life. 2012;64(5):432–442.
- Liu J, et al. Development of novel visual-plus quantitative analysis systems for studying DNA double-strand break repairs in zebrafish. J Genet Genomics. 2012;39(9):489–502.
- Dev H, et al. Shieldin complex promotes DNA end-joining and counters homologous recombination in BRCA1-null cells. Nat Cell Biol. 2018;20(8):954–965.
- Boersma V, et al. MAD2L2 controls DNA repair at telomeres and DNA breaks by inhibiting 5’ end resection. Nature. 2015;521(7553):537–540.
- Iacovoni JS, et al. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 2010;29(8):1446–1457.
- Puram SV, et al. The ubiquitin receptor S5a/Rpn10 links centrosomal proteasomes with dendrite development in the mammalian brain. Cell Rep. 2013;4(1):19–30.
- Wang Q, et al. Structure of S5a bound to monoubiquitin provides a model for polyubiquitin recognition. J Mol Biol. 2005;348(3):727–739.
- Riedinger C, et al. Structure of Rpn10 and its interactions with polyubiquitin chains and the proteasome subunit Rpn12. J Biol Chem. 2010;285(44):33992–34003.
- Mailand N, et al. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell. 2007;131(5):887–900.
- Le Tallec B, et al. Hsm3/S5b participates in the assembly pathway of the 19S regulatory particle of the proteasome. Mol Cell. 2009;33(3):389–399.
- Husnjak K, et al. Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature. 2008;453(7194):481–488.
- Li T, et al. A promiscuous alpha-helical motif anchors viral hijackers and substrate receptors to the CUL4-DDB1 ubiquitin ligase machinery. Nat Struct Mol Biol. 2010;17(1):105–111.
- Fong PC, et al. Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med. 2009;361(17):123–134.
- McCann TS, Tansey WP. Functions of the proteasome on chromatin. Biomolecules. 2014;4(4):1026–1044.
- Ferdous A, et al. The 19S regulatory particle of the proteasome is required for efficient transcription elongation by RNA polymerase II. Mol Cell. 2001;7(5):981–991.
- Seo HD, et al. The 19S proteasome is directly involved in the regulation of heterochromatin spreading in fission yeast. J Biol Chem. 2017;292(41):17144–17155.
- Tomimatsu N, et al. DNA-damage-induced degradation of EXO1 exonuclease limits DNA end resection to ensure accurate DNA repair. J Biol Chem. 2017;292(26):10779–10790.
- Taylor BJ, et al. DNA deaminases induce break-associated mutation showers with implication of APOBEC3B and 3A in breast cancer kataegis. Elife. 2013;2:e00534.
- Burns MB, et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature. 2013;494(7437):366–370.
- Ochs F, et al. 53BP1 fosters fidelity of homology-directed DNA repair. Nat Struct Mol Biol. 2016;23(8):714–721.
- Farmer H, et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature. 2005;434(7035):917–921.
- Harrision D, et al. Poly(ADP-ribose) glycohydrolase (PARG) vs. poly(ADP-ribose) polymerase (PARP) — function in genome maintenance and relevance of inhibitors for anti-cancer therapy. Front Mol Biosci. 2020;7:191. View this article via: CrossRef Google Scholar
- Radnai B, et al. Protective effect of the poly(ADP-ribose) polymerase inhibitor PJ34 on mitochondrial depolarization-mediated cell death in hepatocellular carcinoma cells involves attenuation of c-Jun N-terminal kinase-2 and protein kinase B/Akt activation. Mol Cancer. 2012;11:34.
- Shimizu S, et al. Expression of poly(ADP-ribose) polymerase in human hepatocellular carcinoma and analysis of biopsy specimens obtained under sonographic guidance. Oncol Rep. 2004;12(4):821–825.View this article via: PubMed Google Scholar
- Nomura F, et al. Enhancement of poly-adenosine diphosphate-ribosylation in human hepatocellular carcinoma. J Gastroenterol Hepatol. 2000;15(5):529–535.
- Davies H, et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat Med. 2017;23(4):517–525.
- Mathur D, et al. Analysis of the mouse embryonic stem cell regulatory networks obtained by ChIP-chip and ChIP-PET. Genome Biol. 2008;9(8):R126.
-
Version history
- Version 1 (October 10, 2023): In-Press Preview
- Version 2 (December 1, 2023): Electronic publication



Copyright © 2024 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)