Advertisement
Research ArticleMuscle biologyNeuroscience Free access | 10.1172/JCI99042
Androgen receptor polyglutamine expansion drives age-dependent quality control defects and muscle dysfunction
Samir R. Nath,1,2,3 Zhigang Yu,1 Theresa A. Gipson,4 Gregory B. Marsh,5 Eriko Yoshidome,1 Diane M. Robins,6 Sokol V. Todi,5 David E. Housman,4 and Andrew P. Lieberman1
1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by
Nath, S.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by Yu, Z. in: JCI | PubMed | Google Scholar
1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by Gipson, T. in: JCI | PubMed | Google Scholar
1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by Marsh, G. in: JCI | PubMed | Google Scholar
1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by Yoshidome, E. in: JCI | PubMed | Google Scholar
1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by
Robins, D.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by
Todi, S.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by Housman, D. in: JCI | PubMed | Google Scholar
1Department of Pathology,
2Medical Scientist Training Program, and
3Cellular and Molecular Biology Graduate Program, University of Michigan Medical School, Ann Arbor, Michigan, USA.
4Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
5Department of Pharmacology, Wayne State University School of Medicine, Detroit, Michigan, USA.
6Department of Human Genetics, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
Find articles by Lieberman, A. in: JCI | PubMed | Google Scholar
Published May 29, 2018 - More info
J Clin Invest. 2018;128(8):3630–3641. https://doi.org/10.1172/JCI99042.
Copyright © 2018, American Society for Clinical Investigation
Received: December 5, 2017; Accepted: May 24, 2018
-
Abstract
Skeletal muscle has emerged as a critical, disease-relevant target tissue in spinal and bulbar muscular atrophy, a degenerative disorder of the neuromuscular system caused by a CAG/polyglutamine (polyQ) expansion in the androgen receptor (AR) gene. Here, we used RNA-sequencing (RNA-Seq) to identify pathways that are disrupted in diseased muscle using AR113Q knockin mice. This analysis unexpectedly identified substantially diminished expression of numerous ubiquitin/proteasome pathway genes in AR113Q muscle, encoding approximately 30% of proteasome subunits and 20% of E2 ubiquitin conjugases. These changes were age, hormone, and glutamine length dependent and arose due to a toxic gain of function conferred by the mutation. Moreover, altered gene expression was associated with decreased levels of the proteasome transcription factor NRF1 and its activator DDI2 and resulted in diminished proteasome activity. Ubiquitinated ADRM1 was detected in AR113Q muscle, indicating the occurrence of stalled proteasomes in mutant mice. Finally, diminished expression of Drosophila orthologues of NRF1 or ADRM1 promoted the accumulation of polyQ AR protein and increased toxicity. Collectively, these data indicate that AR113Q muscle develops progressive proteasome dysfunction that leads to the impairment of quality control and the accumulation of polyQ AR protein, key features that contribute to the age-dependent onset and progression of this disorder.
-
Introduction
Degeneration of the neuromuscular system is characteristic of the age-dependent phenotype that manifests in subjects with spinal and bulbar muscular atrophy (SBMA), a CAG/polyglutamine (polyQ) expansion disorder caused by a mutation in the androgen receptor (AR) gene (1–4). The polyQ AR undergoes hormone-dependent nuclear translocation and protein unfolding, steps that are essential to disease pathogenesis. Expansion of the AR’s polyQ tract both impairs its normal function as a transcriptional regulator and leads to ligand-dependent proteotoxicity. These features result in the disruption of multiple pathways that are critical for normal cellular function (5–13).
Among the tissues affected in SBMA, skeletal muscle is a critical target that contributes to disease pathogenesis. This concept is supported by clinical observations and data from experimental models. Subjects with SBMA exhibit evidence of muscle toxicity, including myopathic features on muscle biopsy and elevated serum creatine kinase levels that are higher than normally found in purely denervating diseases (14–16). Moreover, muscle satellite cells cultured from SBMA patients show androgen-dependent impairment of fusion to form myotubes (17), indicating cell-autonomous toxicity in muscle. Supporting the notion that myopathy is an early disease manifestation, knockin mice expressing the polyQ AR develop myopathy months prior to spinal cord pathology (18). Further support for the role of muscle in disease pathogenesis is provided by data from transgenic mice that overexpress the WT AR only in skeletal muscle and show hormone-dependent myopathy and motor axon loss (19); similar effects are seen in transgenic mice overexpressing polyQ AR only in muscle (20). Notably, the phenotypic severity of SBMA transgenic mice is ameliorated by overexpression of insulin-like growth factor-1 in muscle (21). That muscle contributes to the SBMA phenotype and provides a therapeutic target is further corroborated by studies showing that peripheral polyQ AR knockdown by antisense oligonucleotides (ASOs) or conditional gene deletion in skeletal muscle rescues disease in mice (22, 23).
While this body of evidence indicates an important role for skeletal muscle in the pathogenesis of neuromuscular degeneration in SBMA, little is known about the pathways that contribute to this process. Here, we used an unbiased approach to identify gene-expression changes in skeletal muscle from AR113Q mice, a knockin mouse model that displays a hormone-dependent phenotype characterized by diminished body weight, impaired strength, and shortened life span (18). This analysis unexpectedly uncovered decreased expression of numerous genes encoding components of the ubiquitin/proteasome pathway. We demonstrate that these changes are age, hormone, and glutamine tract length dependent, occur in multiple disease-relevant muscles, and are associated with diminished proteasome function. Our analysis identifies progressive impairment of a critical protein quality control pathway in SBMA mice, a feature that likely contributes to the age dependence of this disorder.
-
Results
Age-dependent diminished expression of ubiquitin/proteasome pathway genes in AR113Q skeletal muscle. To identify pathways disrupted by the polyQ AR that contribute to skeletal muscle pathology, we performed RNA-sequencing (RNA-Seq) on the levator ani/bulbocavernosus (LABC) muscle from littermate WT and AR113Q males at 14 weeks of age. The LABC is a pelvic floor muscle that was selected because of its sensitivity to polyQ AR toxicity owing to its high level of polyQ AR expression and aggregation. As such, this muscle is markedly atrophic in sexually mature AR113Q males at a young age (Figure 1A). Our analysis uncovered a large number of gene-expression changes in mutant muscle, including many affecting regulators of cellular energy metabolism, including Gapdh, Hk2, Pgk1, AldoA, and Pgam2 (Supplemental Table 1; supplemental material available online with this article; https://doi.org/10.1172/JCI99042DS1), as has been previously reported (9, 24, 25). Unexpectedly, we also uncovered diminished expression of numerous components of the ubiquitin/proteasome pathway in mutant muscle. These changes involved approximately 30% of constitutive proteasome subunits (P < 0.002 by hypergeometric test) and approximately 20% of E2 ubiquitin–conjugating enzymes (P < 0.05 by hypergeometric test), all of which were reduced in expression in the 14-week LABC from AR113Q males (Figure 1B and Supplemental Tables 1 and 2); many more subunits trended toward reduced expression, but did not reach statistical significance. In contrast, no proteasome subunits were increased in expression, including immunoproteasome subunits and assembly chaperones. Importantly, the observed changes included several key subunits, such as ubiquitin receptors, proteolytic subunits, and assembly scaffold proteins.
 Figure 1
Figure 1Decreased expression of ubiquitin/proteasome pathway genes in AR113Q muscle. (A) LABC muscle from 14-week-old WT or AR113Q (113Q) males stained by FITC–wheat germ agglutinin (WGA) to highlight muscle fibers. Scale bar: 100 μm. Fiber size quantified at right and expressed as relative cross-sectional area (CSA). Data are shown as mean ± SEM. n = 3 mice/genotype. ****P < 0.0001, t test. (B) Diagram of the 26S proteasome, highlighting subunits of the 20S core and the 19S lid. Genes with reduced expression in 14-week LABC of AR113Q vs. WT mice were identified by RNA-Seq and are color coded in blue according to fold change. Those that met a significance threshold of less than 0.67 fold change and an FDR of less than 0.05 are marked with white text labels.
We confirmed in an independent cohort of AR113Q males that proteasome subunits Psma7, a 20S core scaffold protein (26, 27), and Psmd6, a regulatory subunit of the lid (28), and the ubiquitin conjugase Ube2T were reduced in expression in the 14-week LABC (Figure 2). We also aged a cohort of AR113Q and WT littermates to 52 weeks, at which point mutant males exhibited a severe phenotype characterized by significantly diminished body weight, grip strength, muscle fiber size, and limb muscle mass (Supplemental Figure 1). These mice continued to display significantly decreased expression of Psma7, Psmd6, and Ube2T in the LABC (Figure 2). Furthermore, we found that these same gene-expression changes also occurred in other disease-relevant skeletal muscles, including the quadriceps and tibialis anterior, in an age-dependent manner (Figure 2). Notably, these changes were not detected in 52-week lumbar spinal cord, the site that innervates these muscles (Figure 2). Although these data do not preclude the occurrence of altered gene expression in a small subset of cells, such as motor neurons, our findings suggest that the gene-expression changes were most severe in skeletal muscles of AR113Q mice.
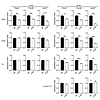 Figure 2
Figure 2Decreased expression of ubiquitin/proteasome pathway genes is age dependent. Relative expression of proteasome genes Psma7 and Psmd6 and the ubiquitin conjugase Ube2T was determined in WT or AR113Q mice at 14 or 52 weeks by qPCR. Shown are data from LABC (n = 3 mice/genotype), quadriceps (Quad) (14 weeks, n = 3 mice/genotype; 52 weeks, n = 6 mice/genotype), tibialis anterior (T.A.) (n = 3 mice/genotype), and lumbar spinal cord (S.C.) (n = 3 mice/genotype). Data are shown as mean ± SEM.*P < 0.05; **P < 0.01; ***P < 0.001, t test.
Gene-expression changes are polyQ length dependent and mediated by a toxic gain of function. Pathologic expansion of the AR’s polyQ tract leads to both a partial loss of normal function and ligand-dependent proteotoxicity. We sought to determine the extent to which these 2 consequences of the mutation contribute to age-dependent loss of ubiquitin/proteasome pathway gene expression. To address this question, we first determined whether changes in gene expression were solely dependent upon the length of the AR’s polyQ tract. To accomplish this, we compared gene expression in 2 lines of knockin mice generated using the same targeting vector except for the length of the CAG repeat (29, 30). Whereas AR113Q mice develop age-dependent neuromuscular pathology, AR21Q males are similar to WT littermates (18). Expression of Psma7, Psmd6, and Ube2T was significantly decreased in 14-week LABC of AR113Q versus AR21Q males, establishing that these changes are dependent upon the length of the polyQ tract (Figure 3A). Glutamine tract length dependence was corroborated in mutant males in which the CAG repeat had contracted to encode 86 glutamines and that develop a more slowly progressive phenotype. These mice showed an intermediate loss of expression in the 26-week quadriceps compared with WT and AR113Q mice (Supplemental Figure 3).
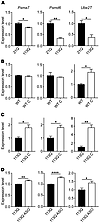 Figure 3
Figure 3Ubiquitin/proteasome pathway gene-expression changes are hormone and glutamine length dependent. (A) Relative expression of Psma7, Psmd6, and Ube2T was determined in LABC from 14-week AR21Q or AR113Q mice by qPCR. Data are shown as mean ± SEM. n = 3 mice/genotype. (B and C) WT (B) and AR113Q (C) males were surgically castrated (C) at 5 to 6 weeks, then aged to 14 weeks. Gene expression in LABC was determined by qPCR and compared with that of noncastrated littermates. Data are shown as mean ± SEM. n = 3 mice/genotype. (D) AR113Q males were treated with AR-targeted ASO or vehicle from 6 to 26 weeks. Gene expression in quadriceps was determined by qPCR. Data are shown as mean ± SEM. n = 3–4 mice/genotype. *P < 0.05; **P < 0.01; ****P < 0.0001, t test.
To probe how polyQ AR alters gene expression, we asked whether these changes represent a loss of normal AR signaling or a gain of toxic function. To answer this question, we first examined the effect of surgical castration in WT males. This intervention leads to loss of testosterone, the ligand for AR, and as a result, eliminates AR signaling. Castration alone had no effect on expression of the proteasome genes Psma7 and Psmd6 and increased expression of Ube2T (Figure 3B), indicating that loss of AR signaling as a result of testosterone removal was not sufficient to account for the diminished gene expression characteristic of AR113Q muscle.
Next, we asked whether these changes represent a toxic gain of function of the disease-causing protein. In contrast to effects in WT males, surgical castration of AR113Q males significantly rescued gene-expression changes (Figure 3C). These findings are consistent with other manifestations of the neuromuscular phenotype of AR113Q males that are dependent upon both polyQ AR and its ligand testosterone (18), reflecting ligand-dependent proteotoxicity. The essential role of polyQ AR expression in muscle to effect these changes was verified by subcutaneous administration of AR-targeted ASOs. Treatment of AR113Q mice with ASOs from age 6 to 26 weeks rescues disease phenotypes and diminishes polyQ AR expression in skeletal muscle, but not spinal cord (22). This treatment also increased expression of Psma7, Psmd6, and Ube2T in quadriceps muscle of mutant males (Figure 3D). Collectively, these data show that diminished expression of ubiquitin/proteasome pathway genes is polyQ length dependent and due to a toxic gain of function rather than loss of AR signaling.
Loss of proteasome subunits is associated with diminished proteasome transcriptional machinery. Coincident with diminished gene expression, we found that AR113Q muscle showed robust and significant reduction in several proteasome subunits when analyzed by Western blot (P < 0.05). Subunits altered in 14-week LABC included PSMB7, the trypsin-like subunit (31), PSMB5, the chymotrypsin-like subunit (31), PSMD11, which functions as a regulatory link between the ATPase ring and 20S core (32), and PSMA7, a scaffold protein and contributor to the size gate of the proteasome (26, 27) (Figure 4A). We further confirmed changes in PSMB5 and PSMB7 protein levels in 52-week quadriceps muscle (Figure 4B). Notably, all proteasome subunits that were found to be downregulated at the protein level also showed reduced mRNA expression, except for PSMB5 (Supplemental Figure 2). The finding that this subunit was decreased at the protein, but not mRNA, level raises the possibility that the level of the entire proteasome machinery may be diminished in mutant muscle.
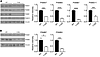 Figure 4
Figure 4Proteasome subunits are decreased in AR113Q muscle. (A and B) LABC (A) was harvested at 14 weeks and quadriceps (B) at 52 weeks from WT and AR113Q mice. Samples were analyzed by Western blot for indicated proteasome subunits. Vinculin controls were used for loading. Quantification is shown at right. Data are shown as mean ± SEM. n = 6 mice/group for PSMA7 (LABC); n = 7 mice/genotype for PSMB5, PSMB7, and PSMD11 (LABC); and n = 3 mice/genotype (quadriceps). *P < 0.05; ***P < 0.001; ****P < 0.0001, t test.
Given the broad loss of proteasome subunit expression, we hypothesized that critical transcriptional activators of proteasome genes are altered in AR113Q muscle. To test this notion, we examined the best-characterized proteasome transcription factor, nuclear factor erythroid 2-like 1 (NRF1/NFE2L1) (33). We found a profound loss of the 95 kDa form of NRF1 in both 14-week LABC (Figure 5A) and 52-week quadriceps muscle (Figure 5B) from AR113Q males. This observation was confirmed using a second anti-NRF1 antibody (Supplemental Figure 4A). Notably, the 95 kDa NRF1 species is the cleaved and active form of this transcription factor (34), and the changes we observed in AR113Q mice occurred in the absence of changes in Nrf1/Nfe2l1 gene expression (Supplemental Figure 4B). This led us to hypothesize that factors that generate the cleaved NRF1 species are lost in disease. Recent work from multiple groups has demonstrated that the mammalian aspartyl protease DDI2 and its Caenorhabditis elegans orthologue SKN-1A cleave NRF1 from the ER membrane to allow nuclear translocation and transcription of proteasomal genes (35, 36). Intriguingly, both mRNA and protein levels of Ddi2 were diminished in the 14-week LABC of AR113Q mice (Figure 5, C and D). Furthermore, the loss of Ddi2 gene expression was rescued by surgical castration, similarly to proteasome gene-expression changes, suggesting that diminished DDI2 may contribute to decreased 95 kDa Nrf1 species in AR113Q muscle.
 Figure 5
Figure 5NRF1 and DDI2 are reduced in AR113Q muscle, and reduction of NRF1 leads to a buildup of polyQ AR in vivo. (A and B) Lysates from 14-week LABC (A) or 52-week quadriceps (B) were probed by Western blot for expression of the proteasome transcription factor NRF1 (NFE2L1). Vinculin controls were used for loading. Quantification is shown at right. Data are shown as mean ± SEM. n = 3 mice/genotype (quadriceps); n = 4 mice/genotype (LABC). **P < 0.01; ****P < 0.0001, t test. (C) Expression of Ddi2 mRNA was quantified in 14-week LABC from WT, AR113Q, and castrated AR113Q mice by qPCR. Data are shown as mean ± SEM. n = 3 mice/genotype. *P < 0.05; **P < 0.01, 1-way ANOVA with Tukey’s post hoc analysis. (D) Lysates from 14-week LABC were probed by Western blot for expression of DDI2. Vinculin controls were used for loading. Quantification is shown at right. Data are shown as mean ± SEM. n = 4 mice/genotype. ****P < 0.0001, t test. (E) Drosophila expressing AR52Q under control of the GMR-gal4 driver were crossed to 3 independent lines expressing RNAi targeted toward cap’n’collar (cnc), the NRF1 orthologue. Levels of AR were analyzed by Western blot. Tubulin controls were used for loading. Relative intensity of high–molecular weight (HMW) AR species is quantified below. Data are shown as mean ± SEM. n = 5 biological replicates/group. *P < 0.05; **P < 0.01, 1-way ANOVA with Tukey’s post hoc analysis. ctrl, control.
To determine whether loss of the proteasome transcriptional machinery is sufficient to alter steady-state levels of polyQ AR, we used a well-characterized Drosophila model expressing AR52Q under the control of the GMR-Gal4 promoter (13). We crossed this model to 3 independent lines expressing RNAi toward cap’n’collar, the fly orthologue of NRF1 (37). Target knockdown was confirmed by quantitative PCR (qPCR) (Supplemental Figure 5A). Cap’n’collar knockdown causes diminished expression of proteasome genes in flies (37). This manipulation resulted in significant accumulation of a high–molecular weight polyQ AR species in 2 of 3 lines, with the third line yielding results that trended similarly (P = 0.06) (Figure 5E). This accumulation of polyQ AR protein was accompanied by evidence of enhanced toxicity (Supplemental Figure 5D). We conclude that loss of NRF1 impairs polyQ AR clearance in vivo and that this accumulation contributes to age-dependent toxicity in vivo.
Proteasome activity is diminished in AR113Q mice. Following our results demonstrating reduced levels of NRF1, DDI2, and proteasome subunits, we hypothesized that AR113Q muscle exhibits decreased proteasome activity. To test the functional status of the proteasome, we employed the activity-based probe Me4BodipyFL-Ahx3Leu3VS. This fluorescent probe binds the active site of catalytic proteasome subunits, allowing detection of changes in constitutive proteasome subunits as well as induction of the immunoproteasome (38, 39). We initially confirmed that signal in this assay is inhibited by MG132, a small molecule that similarly binds catalytic sites to inhibit proteasome activity and thereby competes for binding by the activity-based probe (Supplemental Figure 6). Using this assay, we found relatively little difference in proteasome activity in quadriceps muscle at 14 weeks, but significant age-dependent loss of active PSMB5 and PSMB6/7 bands in 52-week quadriceps of AR113Q mice (Figure 6, A and B). These changes were not associated with the appearance of additional bands that might suggest induction of the immunoproteasome. Corroborating these findings in 52-week quadriceps muscle, we observed a significant reduction in chymotrypsin-like activity of the proteasome, as measured by cleavage of the fluorogenic substrate Suc-AMC-LLVY (Figure 6C).
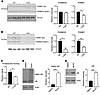 Figure 6
Figure 6Proteasome activity is diminished in AR113Q muscle. (A and B) Lysates from 14-week (A) and 52-week (B) quadriceps were incubated with a BODIPY-labeled activity-based probe, then resolved by SDS-PAGE. Quantification is shown at right. Data are shown as mean ± SEM. n = 3 mice/genotype. *P < 0.05; **P < 0.01, t test. (C) Lysates from quadriceps were prepared from 52-week WT and AR113Q mice and probed for chymotrypsin-like activity of the proteasome by cleavage of AMC-Suc-LLVY. Data are shown as mean ± SEM. n = 5 mice/genotype. *P < 0.05, t test. (D and E) PC12 cells were induced to express AR112Q for 48 hours in the presence of 10 nM R1881, then washed with PBS and incubated with 100 nM epoxomicin (Epox), 1 μM MLN7243 (MLN7), or vehicle (DMSO) for 18 hours, as indicated. AR levels were determined by Western blot. Quantification is shown at right. Data are shown as mean ± SEM. n = 3 biological replicates. ***P < 0.001, t test.
To assess the consequence of diminished functional proteasomes on polyQ AR clearance, we used PC12 cells stably expressing AR112Q under the control of a TetOn promoter (40). Inhibition of the proteasome by epoxomicin led to the accumulation of high–molecular weight polyQ AR (Figure 6D), suggestive of ubiquitinated or aggregated species and paralleling our findings in Drosophila following knockdown of cap’n’collar (Figure 5E). Furthermore, inhibition of ubiquitination by MLN7243, a small molecule inhibitor of E1 ubiquitin–activating enzymes (41), led to accumulation of AR112Q monomer, demonstrating the importance of ubiquitination and an intact proteasome on polyQ AR clearance (Figure 6E).
Proteasome lid modifications suggest impairment of polyQ AR degradation. The importance of ubiquitination for polyQ AR clearance prompted us to examine the proteasome subunits responsible for binding ubiquitinated substrates in AR113Q muscle. Ubiquitination of the proteasome’s ubiquitin receptor ADRM1/Rpn13 has been shown to occur, causing blockade of ubiquitin-dependent degradation and leading to a buildup of “stalled” proteasomes (42). To assess whether this occurs in AR113Q mice, we examined ADRM1 protein in WT and AR113Q 14-week LABC and 52-week quadriceps. Strikingly, we found evidence of high–molecular weight bands and diminished levels of unmodified ADRM1 in these mutant muscles (Figure 7A and Supplemental Figure 7A). To confirm that high–molecular weight ADRM1 bands reflected ubiquitinated species, samples were incubated with the deubiquitinating (DUB) enzyme USP2 in the absence or presence of the DUB inhibitor N-ethylmaleimide (NEM). Treatment with USP2 diminished the amount of high–molecular weight ADRM1 and increased the amount of unmodified ADRM1 only in the absence of NEM (Figure 7B and Supplemental Figure 7B). These data indicate that high–molecular weight ADRM1 species are ubiquitinated and that these ubiquitin chains are cleaved by the active DUB. We further confirmed ADRM1 ubiquitination in AR113Q muscle by ADRM1 IP, followed by Western blot for ubiquitin (Supplemental Figure 7C).
 Figure 7
Figure 7ADRM1 is ubiquitinated in AR113Q mouse muscle. (A) Lysates of 52-week quadriceps were probed by Western blot for ADRM1. Vinculin controls were used for loading. Quantification is shown at right. Data are shown as mean ± SEM. n = 3 mice/genotype. **P < 0.01; ***P < 0.001, t test. (B) Lysates of 52-week quadriceps were incubated with the catalytic domain of USP2 in the absence or presence of 10 μM NEM, as indicated, and then analyzed by Western blot. High–molecular weight species were normalized to level of ADRM1 monomer. Quantification is shown at right. n = 3 mice/genotype. *P < 0.05, t test. (C) Drosophila expressing AR52Q under control of the GMR-Gal4 driver were crossed to lines expressing RNAi against Rpn11 or Rpn13/Adrm1. Levels of AR were analyzed by Western blot. Tubulin controls were used for loading. Relative intensity of high–molecular weight AR species is quantified at right. *P < 0.05; **P < 0.01, 1-way ANOVA with Tukey’s post hoc analysis. (D) PC12 cells were induced to express AR112Q for 72 hours in the presence of 10 nM R1881, then washed with PBS and treated with DMSO or 3 μM RA190 to inhibit Rpn13/ADRM1 for 18 hours. Data are shown as mean ± SEM. n = 3 biological replicates. **P < 0.01, t test.
To assess the functional importance of this proteasome subunit in polyQ AR clearance, we used Drosophila expressing AR52Q under the control of the GMR-Gal4 promoter. We crossed this line to flies expressing RNAi toward Rpn13/ADRM1. Flies expressing RNAi toward Rpn11, a well-characterized DUB associated with the proteasome that removes ubiquitin chains and allows for substrate entrance (43), were used as a positive control. Target knockdown was confirmed by qPCR (Supplemental Figure 5, B and C). We found that knockdown of either Rpn13/ADRM1 or Rpn11 led to an accumulation of high–molecular weight polyQ AR in vivo (Figure 7C), similarly to the effects of cap’n’collar knockdown (Figure 5E). This effect was accompanied by evidence of enhanced toxicity (Supplemental Figure 5D). Further, treatment of PC12 cells expressing AR112Q with RA190, a small molecule that inhibits ADRM1/Rpn13 (44), similarly yielded an accumulation of high–molecular weight polyQ AR species (Figure 7D), emphasizing the role of the proteasome lid in polyQ AR degradation. Collectively, our results demonstrate that impairment of the proteasome occurs by multiple mechanisms in vivo, including transcriptional silencing and posttranslational modification, and that the loss of functional proteasomes leads to the accumulation of the toxic, mutant protein.
-
Discussion
Recent studies have demonstrated a key role for skeletal muscle in the pathogenesis of the polyQ expansion disease SBMA (4, 22, 23), but the downstream mediators of age- and tissue-dependent decline are not well characterized. This study reports for what we believe is the first time the unexpected finding of proteasome dysfunction in the skeletal muscle from a knockin mouse model of SBMA. Proteasome impairment occurred in an age-, hormone-, and polyQ length–dependent manner in multiple muscles of AR113Q mice. These changes are particularly relevant, as the proteasome is the primary pathway by which both WT and polyQ AR are degraded (6, 45–47). Furthermore, promoting polyQ AR degradation through the proteasome shows phenotypic rescue in mice and is an active area of therapeutic investigation (48–54).
The occurrence of proteasome dysfunction in AR113Q muscle was unanticipated, as skeletal muscle atrophy that follows other injuries, including surgical denervation, fasting, tumor bearing, uremia, and diabetes, is associated with a significant upregulation of proteasome subunits (55). Notably, SBMA is distinguished from these other states by the accumulation of a toxic, misfolded protein in diseased muscle. That the impairment of proteasome function is closely tied to AR113Q protein misfolding is demonstrated by the observation that gene-expression changes occur as a consequence of a gain of function conferred by the polyQ tract expansion (Figure 3). These gene-expression changes affect approximately 30% of proteasome subunits and approximately 20% of E2 ubiquitin–conjugating enzymes. They result in diminished functionally active proteasomes (Figure 6) and an accumulation of stalled proteasomes characterized by the ubiquitination of ADRM1 (Figure 7), a proteasome subunit responsible for binding ubiquitinated substrates. We demonstrate that changes in proteasome function impair clearance of the polyQ AR in well-established cellular and Drosophila models of SBMA (Figures 5 and 7), indicating the importance of quality control defects to disease. Dysregulation of the ubiquitin-proteasome system was of particular interest, since prior studies have established that quality control decisions regulating degradation of the polyQ AR are tightly controlled by the Hsp90/Hsp70-based chaperone machinery (56, 57). Importantly, targeting this pathway promotes clearance of the polyQ AR by the proteasome and alleviates the disease phenotype in SBMA models (48–50, 54, 58, 59). Our findings establish that proteasome function diminishes with age in multiple disease-relevant skeletal muscles from AR113Q mice. This identifies a progressive impairment of the proteostasis machinery that likely contributes to the age-dependent phenotype characteristic of this disorder. We speculate that these changes result in enhanced proteotoxicity, leading to muscle atrophy through distinct and currently poorly characterized mechanisms.
The extent to which proteasome dysfunction occurs in the polyQ diseases has been controversial, as previous findings in vivo have lacked consistency and reproducibility (60). In SBMA, proteasome activity was previously studied in transgenic mice overexpressing AR97Q. While accumulation of the proteasome reporter substrate GFP-Ub was noted, this finding was attributed to increased GFP-Ub gene expression as measured relative to the loading control GAPDH (58). However, recent studies have demonstrated marked downregulation of glycolytic enzymes, including GAPDH, in muscle from SBMA mice and patients as a component of a previously unappreciated metabolic shift (24, 25). Our current analysis takes into account these changes and demonstrates diminished proteasome mRNA and protein levels along with impaired function in AR113Q muscle. Notably, prior studies have also demonstrated activation of macroautophagy in AR113Q and SBMA patient muscle, including increased levels of LC3-II and p62 and increased LC3 puncta (61, 62). These changes may be a compensatory response that is aimed at maintaining organ system homeostasis in the setting of diminished proteasome function.
The consequences of expansion of the AR’s polyQ tract are complex, with a resulting partial loss of function impairing transcriptional regulation of AR target genes and ligand-dependent proteotoxicity disrupting several critical downstream pathways (1–4). Both of these effects are likely to contribute to the SBMA phenotype. In fact, restoring transcriptional regulatory activity to the polyQ AR by disrupting its SUMOylation (AR113Q KRKR mice) rescues exercise capacity and extends life span without altering protein aggregation or diminishing muscle atrophy (63). In contrast, the effects on proteasome function characterized here are independent of the partial loss of function conferred by the polyQ tract expansion. This conclusion is supported by the fact that the changes in ubiquitin/proteasome pathway genes in AR113Q mice persist in skeletal muscle from AR113Q KRKR mutants (data not shown). This gain-of-function effect may be due to either a new, ligand-dependent function of the polyQ protein or an increased, toxic interaction with normal binding partners. More work is warranted to determine the exact link between polyQ AR and loss of NRF1 function.
Collectively, our data demonstrate that the proteasome is impaired in AR113Q muscles by a multifactorial process, including loss of the transcriptional machinery that promotes gene expression and the occurrence of posttranslational modifications to proteasome subunits that impair binding of ubiquitinated substrates. These findings represent the first demonstration, to our knowledge, of DDI2 and ADRM1 alterations in a mouse model of disease and validate the functional relevance of these changes in the context of proteotoxic stress. While the proteasome shows functional impairment in AR113Q muscle, we speculate that enhancing polyQ AR degradation by the proteasome remains a viable therapeutic strategy. Supporting this notion are several reports leveraging the chaperone machinery to achieve this result (48–50, 52, 53, 56, 57, 64–68). We propose that alleviating proteasome dysfunction may confer a synergistic rescue with efforts to target the chaperone machinery. This concept is supported by a recent study demonstrating that activation of the transcription factors NRF1 or NRF2 by a small molecule leads to significant phenotypic rescue in a transgenic mouse model of SBMA (69). By combining strategies such as this with proteasome activation (70), chaperone modulation (48–54), and/or direct manipulation of the protein’s interactors (13, 71), it may be possible to therapeutically alleviate toxic effects of the mutant polyQ protein.
-
Methods
Antibodies and reagents. The following primary antibodies were used in our studies: rabbit anti-AR (catalog sc-7305, Santa Cruz Biotechnology Inc.), mouse anti–β-actin (catalog A5441, MilliporeSigma), mouse anti-vinculin (catalog V9131, MilliporeSigma), rabbit anti-PSMA7 (catalog ab133502, Abcam), rabbit anti-PSMB7 (catalog 14727-1-AP, Protein Tech), rabbit anti-PSMB5 (catalog ab3330, Abcam), rabbit anti-PSMD11 (catalog NBP1-46191, Novus), rabbit anti-ADRM1 (Western blots: catalog ENZ-ABS296-0100, Enzo Life Sciences), rabbit anti-ADRM1 (IP: catalog A302-554A, Bethyl Labs), rabbit anti-Nrf1 (catalog sc-721, Santa Cruz Biotechnology Inc.), mouse anti-Nrf1 (catalog sc-13031, Santa Cruz Biotechnology Inc.), wheat germ agglutinin Alexa Fluor 488 conjugate (catalog W11261, Thermo Fisher Scientific), mouse anti-tubulin (catalog T5168, MilliporeSigma), and DDI2 (provided by Shigeo Murata, University of Tokyo, Tokyo, Japan). The following drugs and small molecules were used: MLN7243 (CT-M7243, Chemietek), RA190 (5.30341.0001, Calbiochem), epoxomicin (E3652, MilliporeSigma), MG132 (M7449, MilliporeSigma), Me4BodipyFL-Ahx3Leu3VS (I-190, Boston Biochem), and Suc-LLVY-AMC (94367-21-2, Enzo Life Sciences).
Mice. AR113Q and AR21 knockin mice were derived using exon 1–specific targeting, as previously described (18, 29, 30), and backcrossed to C57BL/6 (≥10 generations). Mice were housed in a specific pathogen–free facility and maintained on a 12-hour light/12-hour dark cycle with chow and water ad libitum. Genotyping was performed by PCR on ear samples, and primer sequences were as follows: AR forward: 5′-CCAGAATCTGTTCCAGAGCGTG-3′ (MilliporeSigma, 6-FAM labeled); AR reverse: 5′-TGTTCCCCTGGACTCAGATG-3′ (Invitrogen). For ASO treatment, mice were injected from 6 weeks of age to 26 weeks of age once per week subcutaneously with an ASO (50 mg/kg body weight) containing the following sequence: AAGTTGTAGTAGTCGC, as previously described (22). This 16-mer, 2′,4′-constrained ethyl gapmer ASO is complementary to human and mouse AR transcripts.
Orchiectomy. Mice were surgically castrated at 5 to 6 weeks of age according to previously described methods (63). Briefly, a 5 mm incision was made in the abdomen at the level of the hind legs. The vas deferens and spermatic cord were separated and ligated with nonabsorbable sutures. Following ligation, the testes were removed and the incision was closed using an absorbable suture. Mice were allowed to age to 14 weeks following castration, at which time they were euthanized for further analysis.
RNA-Seq. Total RNA was extracted using TRIzol extraction (Ambion) according to the manufacturer’s instructions. cDNA was prepared for sequencing using the TruSeq mRNA-Seq Sample Prep Kit (Illumina) according to the manufacturer’s protocols. Libraries were size selected at 300 bp and sequenced on an Illumina NextSeq platform using the manufacturer’s protocols for paired-end, 80 nt sequencing. Samples were multiplexed into pools of 6 samples each, and each pool was sequenced on 1 lane of the sequencer. The software package Tuxedo Suite (Illumina) was used for alignment, differential expression analysis, and postanalysis diagnostics. The reference transcriptome UCSC mm10 was used for alignment. Expression quantitation and differential expression analysis were performed using Cuffdiff (Gene Pattern). Gene and transcripts were identified as differentially expressed based on 3 criteria: test status was OK, FDR less than 0.05, and fold change 1.5 or greater. Analysis was performed by the University of Michigan Bioinformatics Core. All original RNA-Seq data were deposited in the NCBI’s Gene Expression Omnibus database (GEO GSE106521). Further analysis and visual representation of differentially expressed genes to identify significantly enriched functional categories were performed using GO-Term analysis (Gene Ontology Consortium) and visualized using PantherDB (Bioconductor) (not shown).
qPCR. RNA was isolated from mouse tissues at the indicated time points using a TRIzol (MilliporeSigma) extraction according to the manufacturer’s instructions. RNA (2 μg) was reverse transcribed using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems). qPCR was performed using FastStart TaqMan Probe Master Mix (Roche), a 7500 Real-Time PCR SDS System (Applied Biosystems), and gene-specific FAM-labeled TaqMan primer/probe mixes (Applied Biosystems). Gene expression was normalized to ZFP146-Vic or Cpsf2-Vic multiplexed within the same well.
Protein isolation and Western blot. Indicated tissue or cells were digested in RIPA buffer (Teknova) containing 1 cOmplete Mini Protease Inhibitor Cocktail (Roche) and 5 mM NEM (MilliporeSigma). Tissue was lysed by mechanical homogenization, and cells were lysed by sonication. Muscle lysates were incubated on a rocker at 4°C for 1 hour to allow for complete digestion, then cleared by spinning at 12,000 g for 10 minutes at 4°C. Cell lysates were cleared by spinning at 15,000 g for 15 minutes at 4°C. Protein concentration was calculated using a DC assay (Bio-Rad). Equal amounts of protein were loaded into NuPAGE 4%–12%, 10-well gels (Invitrogen) and run in 1× MOPS buffer (Invitrogen). Gels were transferred to PVDF membranes using a Semi-Dry Transfer System (Bio-Rad). Blots were blocked for 1 hour in 5% nonfat dry milk in TBS containing 0.1% Tween and placed into primary antibodies in blocking solution at 4°C overnight. Goat anti-rabbit HRP (Bio-Rad) and goat anti-mouse HRP (Bio-Rad) were diluted at 1:5,000 in 5% nonfat dried milk and incubated for 1 hour at room temperature. Blots were quantified using ImageStudio Lite (Licor).
IP. For IP, 1.5 mg protein in RIPA buffer (Teknova) containing protease inhibitor cocktail (Roche) and 5 mM NEM (MilliporeSigma) were precleared in 20 μl protein A/G beads for 1 hour at 4°C. Beads were pelleted at 3,000 g for 5 minutes, and lysates were incubated with 22.5 μg of either ADRM1 (Bethyl Labs) antibody or control IgG on a tumbler overnight at 4°C. Protein A beads (20 μl) were then added, and lysates were tumbled at 4°C for 1 hour. Beads were then placed into spin columns (Pierce) and washed 6 times in RIPA. Finally, beads were boiled at 95°C for 10 minutes in ×4 NuPAGE loading buffer (Invitrogen) containing 150 mM DTT and loaded on NuPAGE 12%, 10 well gels (Invitrogen).
Suc-LLVY-AMC fluorogenic cleavage assay. To assess proteasome activity, tissue was lysed by steel bead homogenization in Proteasome Activity Buffer (50 mM HEPES pH 7.5, 20 mM KCl, 5 mM MgCl2, 1 mM DTT, 28 μM ATP) and assayed at 37°C for 40 minutes in the presence or absence of 25 μM MG132. Activity not inhibited by MG132 was considered nonspecific and was subtracted from the uninhibited values.
Activity-based probe assay. To asses levels of active proteasomes, Me4BodipyFL-Ahx3Leu3VS was used according to previously described methods (72). Briefly, tissue was isolated and homogenized in HR buffer (50 mM Tris–HCl [pH 7.4], 5 mM MgCl2, 250 mM sucrose, 1 mM DTT, 2 mM ATP). Protein concentration was determined using the DC assay (Bio-Rad). Protein (40 μg) was incubated with 1 μM Me4BodipyFL-Ahx3Leu3VS for 1 hour at 37°C. Samples were run on 12% NuPAGE Bis-Tris gels (Invitrogen, Thermo Fisher Scientific) at 140V for 2 hours and imaged using a Typhoon Trio Plus Scanner (Amersham Bioscience).
ADRM1 deubiquitination. To reduce ubiquitinated ADRM1, lysate was generated from muscle in cold HR buffer. Tissue was lysed by steel bead homogenization, and 80 μg protein was incubated with 1 μM catalytic domain of USP2 (Boston Biochem) in the presence or absence of 10 μM NEM (MilliporeSigma) at 37°C for 1 hour. Lysates were then run on a 12% NuPAGE Bis-Tris gel and analyzed by Western blot.
Skeletal muscle imaging. Muscle was placed in a vessel containing OCT and frozen in methylbutane cooled by dry ice until solid. Tissue was mounted and cryosectioned using a Cryocut 1800 Cryostat (Leica Biosystems) at 20 μm thickness, rinsed in PBS 3 times, and fixed in ice-cold methanol for 4 minutes. Tissue sections were again washed with PBS 3 times and covered in 10 μg/ml wheat germ agglutinin for 10 minutes at room temperature. Sections were washed 3 additional times with PBS and mounted using Vectashield Mounting Media with DAPI (Vector Laboratories). Sections were imaged on a Nikon A-1 confocal microscope.
AR112Q-expressing PC12 cells. To coat plates for cellular experiments with poly-d-lysine (Millipore), the poly-d-lysine was diluted to 100 μg/ml and placed into each well to allow thorough coverage of the bottom surface. Plates were then incubated at 37°C for 1 hour and subsequently washed with PBS prior to plating. PC12 cells expressing AR10Q or AR112Q under control of the tetOn promotor were used, as previously described (40) (provided by Diane Merry, Thomas Jefferson University, Philadelphia, Pennsylvania, USA). Briefly, cells were grown in media containing 10% charcoal-stripped horse serum (Atlanta Biologicals) and 5% charcoal-stripped FBS (Atlanta Biologicals), 1% penicillin-streptomycin solution (Gibco, Thermo Fisher Scientific), 0.2% hygromycin, and 0.1% geneticin.
Drosophila stocks. Fly crosses were conducted at 25°C and approximately 60% humidity in a controlled diurnal environment. The RNAi lines used were from the Bloomington Drosophila Stock Center and were as follows: isogenic host strain for attP2 (catalog 36303), attP40 (catalog 36304), RPN 11 (catalog 33662), RPN 13 (catalog 42785), and cnc (catalog 25984, 40854, and 32863). The UAS-ARQ52 transgenic line was a gift of J. Paul Taylor, St. Jude’s Children’s Research Hospital (Memphis, Tennessee, USA). GMR-Gal4 was original Bloomington Drosophila Stock #8121.
Fly lysis and Western blotting. Fly heads were dissected from offspring 1 day after eclosing from their pupal cases and processed for Western blots, as previously described (73–75). Heads were collected in sets of 10, then mechanically homogenized in boiling SDS lysis buffer (50 mM Tris pH 6.8, 2% SDS, 10% glycerol, and 100 mM dithiothreitol), sonicated for 15 seconds, boiled for 10 minutes, and centrifuged at 20,000 g at room temperature for 10 minutes. Western blots were developed and quantified using a CCD-equipped VersaDoc 5000MP system and Quantity One software (Bio-Rad).
For further information, see Supplemental Methods.
Statistics. Graphpad Prism 7.0 was used for statistical analysis, with a minimum cutoff of P < 0.05. Unpaired 2-tailed Student’s t test or 1-way ANOVA with Tukey’s post hoc analysis was used as indicated in the figure legends. Outlier analysis was run on each group, and outliers with a P value of less than 0.05 were excluded from analysis. P < 0.05 was considered significant for all statistical tests.
Study approval. All procedures involving mice were approved by the University of Michigan Committee on the Use and Care of Animals (PRO00006114) and conducted in accordance with institutional and federal guidelines.
-
Author contributions
SRN, ZY, TAG, EY, GBM, and SVT performed experiments. SRN, ZY, GBM, EY, SVT, and APL planned experiments and interpreted data. DMR provided a critical reagent. SRN and APL wrote the manuscript. SRN, TAG, DMR, SVT, DEH, and APL edited the manuscript.
-
Acknowledgments
We thank Shigeo Murata for providing the antibody against DDI2 (36). We thank the Massachusetts Institute of Technology BioMicro Center for sequencing of RNA-Seq libraries. We also thank Richard McEachin and the University of Michigan Bioinformatics Core for their analysis of RNA-Seq data. This work was supported by the NIH (R01 NS055746 to APL; T32 GM007863 and T32 GM007315 to SRN; R01 NS086778 to SVT; P30 CA14051 to the Koch Institute) and the University of Michigan Protein Folding Disease Initiative.
Address correspondence to: Andrew P. Lieberman, Department of Pathology, University of Michigan Medical School, 3510 MSRB1, 1150 W. Medical Center Drive, Ann Arbor, Michigan 48109, USA. Phone: 734.647.4624; Email: liebermn@umich.edu.
-
Footnotes
EY’s present address is: Division of Neuroanatomy, Department of Neuroscience, Yamaguchi University School of Medicine, Ube, Yamaguchi, Japan.
Conflict of interest: The authors have declared that no conflict of interest exists.
Reference information: J Clin Invest. 2018;128(8):3630–3641.https://doi.org/10.1172/JCI99042.
-
References
- Kennedy WR, Alter M, Sung JH. Progressive proximal spinal and bulbar muscular atrophy of late onset. A sex-linked recessive trait. Neurology. 1968;18(7):671–680.
- Sobue G, Hashizume Y, Mukai E, Hirayama M, Mitsuma T, Takahashi A. X-linked recessive bulbospinal neuronopathy. A clinicopathological study. Brain. 1989;112(Pt 1):209–232.View this article via: PubMed Google Scholar
- La Spada AR, Wilson EM, Lubahn DB, Harding AE, Fischbeck KH. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature. 1991;352(6330):77–79.
- Giorgetti E, Lieberman AP. Polyglutamine androgen receptor-mediated neuromuscular disease. Cell Mol Life Sci. 2016;73(21):3991–3999.
- Mhatre AN, et al. Reduced transcriptional regulatory competence of the androgen receptor in X-linked spinal and bulbar muscular atrophy. Nat Genet. 1993;5(2):184–188.
- Lieberman AP, Harmison G, Strand AD, Olson JM, Fischbeck KH. Altered transcriptional regulation in cells expressing the expanded polyglutamine androgen receptor. Hum Mol Genet. 2002;11(17):1967–1976.
- Morfini G, Pigino G, Szebenyi G, You Y, Pollema S, Brady ST. JNK mediates pathogenic effects of polyglutamine-expanded androgen receptor on fast axonal transport. Nat Neurosci. 2006;9(7):907–916.
- Szebenyi G, et al. Neuropathogenic forms of huntingtin and androgen receptor inhibit fast axonal transport. Neuron. 2003;40(1):41–52.
- Ranganathan S, Harmison GG, Meyertholen K, Pennuto M, Burnett BG, Fischbeck KH. Mitochondrial abnormalities in spinal and bulbar muscular atrophy. Hum Mol Genet. 2009;18(1):27–42.
- McCampbell A, et al. CREB-binding protein sequestration by expanded polyglutamine. Hum Mol Genet. 2000;9(14):2197–2202.
- Kemp MQ, et al. Impaired motoneuronal retrograde transport in two models of SBMA implicates two sites of androgen action. Hum Mol Genet. 2011;20(22):4475–4490.
- Montie HL, et al. Cytoplasmic retention of polyglutamine-expanded androgen receptor ameliorates disease via autophagy in a mouse model of spinal and bulbar muscular atrophy. Hum Mol Genet. 2009;18(11):1937–1950.
- Nedelsky NB, et al. Native functions of the androgen receptor are essential to pathogenesis in a Drosophila model of spinobulbar muscular atrophy. Neuron. 2010;67(6):936–952.
- Atsuta N, et al. Natural history of spinal and bulbar muscular atrophy (SBMA): a study of 223 Japanese patients. Brain. 2006;129(Pt 6):1446–1455.View this article via: PubMed Google Scholar
- Sorarù G, et al. Spinal and bulbar muscular atrophy: skeletal muscle pathology in male patients and heterozygous females. J Neurol Sci. 2008;264(1-2):100–105.
- Rhodes LE, et al. Clinical features of spinal and bulbar muscular atrophy. Brain. 2009;132(Pt 12):3242–3251.View this article via: PubMed Google Scholar
- Malena A, et al. Androgen-dependent impairment of myogenesis in spinal and bulbar muscular atrophy. Acta Neuropathol. 2013;126(1):109–121.
- Yu Z, et al. Androgen-dependent pathology demonstrates myopathic contribution to the Kennedy disease phenotype in a mouse knock-in model. J Clin Invest. 2006;116(10):2663–2672.
- Monks DA, et al. Overexpression of wild-type androgen receptor in muscle recapitulates polyglutamine disease. Proc Natl Acad Sci U S A. 2007;104(46):18259–18264.
- Ramzan F, et al. Distinct etiological roles for myocytes and motor neurons in a mouse model of Kennedy’s disease/spinobulbar muscular atrophy. J Neurosci. 2015;35(16):6444–6451.
- Palazzolo I, et al. Overexpression of IGF-1 in muscle attenuates disease in a mouse model of spinal and bulbar muscular atrophy. Neuron. 2009;63(3):316–328.
- Lieberman AP, et al. Peripheral androgen receptor gene suppression rescues disease in mouse models of spinal and bulbar muscular atrophy. Cell Rep. 2014;7(3):774–784.
- Cortes CJ, et al. Muscle expression of mutant androgen receptor accounts for systemic and motor neuron disease phenotypes in spinal and bulbar muscular atrophy. Neuron. 2014;82(2):295–307.View this article via: PubMed Google Scholar
- Giorgetti E, et al. Rescue of metabolic slterations in AR113Q skeletal muscle by peripheral androgen receptor gene silencing. Cell Rep. 2016;17(1):125–136.
- Rocchi A, et al. Glycolytic-to-oxidative fiber-type switch and mTOR signaling activation are early-onset features of SBMA muscle modified by high-fat diet. Acta Neuropathol. 2016;132(1):127–144.
- Li D, et al. c-Abl regulates proteasome abundance by controlling the ubiquitin-proteasomal degradation of PSMA7 subunit. Cell Rep. 2015;10(4):484–496.
- Liu X, et al. Interaction between c-Abl and Arg tyrosine kinases and proteasome subunit PSMA7 regulates proteasome degradation. Mol Cell. 2006;22(3):317–327.
- Isono E, Saeki Y, Yokosawa H, Toh-e A. Rpn7 Is required for the structural integrity of the 26 S proteasome of Saccharomyces cerevisiae. J Biol Chem. 2004;279(26):27168–27176.
- Yu Z, et al. Abnormalities of germ cell maturation and sertoli cell cytoskeleton in androgen receptor 113 CAG knock-in mice reveal toxic effects of the mutant protein. Am J Pathol. 2006;168(1):195–204.
- Albertelli MA, Scheller A, Brogley M, Robins DM. Replacing the mouse androgen receptor with human alleles demonstrates glutamine tract length-dependent effects on physiology and tumorigenesis in mice. Mol Endocrinol. 2006;20(6):1248–1260.
- Collins GA, Goldberg AL. The logic of the 26S proteasome. Cell. 2017;169(5):792–806.
- Lokireddy S, Kukushkin NV, Goldberg AL. cAMP-induced phosphorylation of 26S proteasomes on Rpn6/PSMD11 enhances their activity and the degradation of misfolded proteins. Proc Natl Acad Sci U S A. 2015;112(52):E7176–E7185.
- Steffen J, Seeger M, Koch A, Krüger E. Proteasomal degradation is transcriptionally controlled by TCF11 via an ERAD-dependent feedback loop. Mol Cell. 2010;40(1):147–158.
- Sha Z, Goldberg AL. Reply to Vangala et al.: Complete inhibition of the proteasome reduces new proteasome production by causing Nrf1 aggregation. Curr Biol. 2016;26(18):R836–R837.
- Lehrbach NJ, Ruvkun G. Proteasome dysfunction triggers activation of SKN-1A/Nrf1 by the aspartic protease DDI-1. Elife. 2016;5:e17721. View this article via: PubMed Google Scholar
- Koizumi S, et al. The aspartyl protease DDI2 activates Nrf1 to compensate for proteasome dysfunction. Elife. 2016;5:e18357. View this article via: PubMed Google Scholar
- Grimberg KB, Beskow A, Lundin D, Davis MM, Young P. Basic leucine zipper protein Cnc-C is a substrate and transcriptional regulator of the Drosophila 26S proteasome. Mol Cell Biol. 2011;31(4):897–909.
- Verdoes M, et al. A fluorescent broad-spectrum proteasome inhibitor for labeling proteasomes in vitro and in vivo. Chem Biol. 2006;13(11):1217–1226.
- Carmony KC, Kim KB. Activity-based imaging probes of the proteasome. Cell Biochem Biophys. 2013;67(1):91–101.
- Walcott JL, Merry DE. Ligand promotes intranuclear inclusions in a novel cell model of spinal and bulbar muscular atrophy. J Biol Chem. 2002;277(52):50855–50859.
- Kuo CL, Goldberg AL. Ubiquitinated proteins promote the association of proteasomes with the deubiquitinating enzyme Usp14 and the ubiquitin ligase Ube3c. Proc Natl Acad Sci U S A. 2017;114(17):E3404–E3413.
- Besche HC, et al. Autoubiquitination of the 26S proteasome on Rpn13 regulates breakdown of ubiquitin conjugates. EMBO J. 2014;33(10):1159–1176.
- Ristic G, Tsou WL, Todi SV. An optimal ubiquitin-proteasome pathway in the nervous system: the role of deubiquitinating enzymes. Front Mol Neurosci. 2014;7:72. View this article via: PubMed Google Scholar
- Anchoori RK, et al. A bis-benzylidine piperidone targeting proteasome ubiquitin receptor RPN13/ADRM1 as a therapy for cancer. Cancer Cell. 2013;24(6):791–805.
- Cardozo CP, et al. C-terminal Hsp-interacting protein slows androgen receptor synthesis and reduces its rate of degradation. Arch Biochem Biophys. 2003;410(1):134–140.
- Bailey CK, Andriola IF, Kampinga HH, Merry DE. Molecular chaperones enhance the degradation of expanded polyglutamine repeat androgen receptor in a cellular model of spinal and bulbar muscular atrophy. Hum Mol Genet. 2002;11(5):515–523.
- Segnitz B, Gehring U. The function of steroid hormone receptors is inhibited by the hsp90-specific compound geldanamycin. J Biol Chem. 1997;272(30):18694–18701.
- Wang AM, et al. Activation of Hsp70 reduces neurotoxicity by promoting polyglutamine protein degradation. Nat Chem Biol. 2013;9(2):112–118.
- Adachi H, et al. Heat shock protein 70 chaperone overexpression ameliorates phenotypes of the spinal and bulbar muscular atrophy transgenic mouse model by reducing nuclear-localized mutant androgen receptor protein. J Neurosci. 2003;23(6):2203–2211.
- Adachi H, et al. CHIP overexpression reduces mutant androgen receptor protein and ameliorates phenotypes of the spinal and bulbar muscular atrophy transgenic mouse model. J Neurosci. 2007;27(19):5115–5126.
- Cummings CJ, Mancini MA, Antalffy B, DeFranco DB, Orr HT, Zoghbi HY. Chaperone suppression of aggregation and altered subcellular proteasome localization imply protein misfolding in SCA1. Nat Genet. 1998;19(2):148–154.
- Cummings CJ, et al. Over-expression of inducible HSP70 chaperone suppresses neuropathology and improves motor function in SCA1 mice. Hum Mol Genet. 2001;10(14):1511–1518.
- Miller VM, et al. CHIP suppresses polyglutamine aggregation and toxicity in vitro and in vivo. J Neurosci. 2005;25(40):9152–9161.
- Waza M, et al. 17-AAG, an Hsp90 inhibitor, ameliorates polyglutamine-mediated motor neuron degeneration. Nat Med. 2005;11(10):1088–1095.
- Sacheck JM, et al. Rapid disuse and denervation atrophy involve transcriptional changes similar to those of muscle wasting during systemic diseases. FASEB J. 2007;21(1):140–155.
- Pratt WB, Gestwicki JE, Osawa Y, Lieberman AP. Targeting Hsp90/Hsp70-based protein quality control for treatment of adult onset neurodegenerative diseases. Annu Rev Pharmacol Toxicol. 2015;55:353–371.
- Pratt WB, Morishima Y, Gestwicki JE, Lieberman AP, Osawa Y. A model in which heat shock protein 90 targets protein-folding clefts: rationale for a new approach to neuroprotective treatment of protein folding diseases. Exp Biol Med (Maywood). 2014;239(11):1405–1413.
- Tokui K, et al. 17-DMAG ameliorates polyglutamine-mediated motor neuron degeneration through well-preserved proteasome function in an SBMA model mouse. Hum Mol Genet. 2009;18(5):898–910.
- Waza M, et al. Modulation of Hsp90 function in neurodegenerative disorders: a molecular-targeted therapy against disease-causing protein. J Mol Med. 2006;84(8):635–646.
- Nath SR, Lieberman AP. The ubiquitination, disaggregation and proteasomal degradation machineries in polyglutamine disease. Front Mol Neurosci. 2017;10:78. View this article via: PubMed Google Scholar
- Rusmini P, et al. Aberrant autophagic response in the muscle of a knock-in mouse model of spinal and bulbar muscular atrophy. Sci Rep. 2015;5:15174. View this article via: PubMed Google Scholar
- Chua JP, et al. Transcriptional activation of TFEB/ZKSCAN3 target genes underlies enhanced autophagy in spinobulbar muscular atrophy. Hum Mol Genet. 2014;23(5):1376–1386.
- Chua JP, et al. Disrupting SUMOylation enhances transcriptional function and ameliorates polyglutamine androgen receptor-mediated disease. J Clin Invest. 2015;125(2):831–845.
- Al-Ramahi I, et al. CHIP protects from the neurotoxicity of expanded and wild-type ataxin-1 and promotes their ubiquitination and degradation. J Biol Chem. 2006;281(36):26714–26724.
- Glover JR, Lindquist S. Hsp104, Hsp70, and Hsp40: a novel chaperone system that rescues previously aggregated proteins. Cell. 1998;94(1):73–82.
- Hjerpe R, et al. UBQLN2 mediates autophagy-independent protein aggregate clearance by the proteasome. Cell. 2016;166(4):935–949.
- Jana NR, et al. Co-chaperone CHIP associates with expanded polyglutamine protein and promotes their degradation by proteasomes. J Biol Chem. 2005;280(12):11635–11640.
- Jinwal UK, et al. Chemical manipulation of hsp70 ATPase activity regulates tau stability. J Neurosci. 2009;29(39):12079–12088.
- Bott LC, et al. A small-molecule Nrf1 and Nrf2 activator mitigates polyglutamine toxicity in spinal and bulbar muscular atrophy. Hum Mol Genet. 2016;25(10):1979–1989.
- Myeku N, et al. Tau-driven 26S proteasome impairment and cognitive dysfunction can be prevented early in disease by activating cAMP-PKA signaling. Nat Med. 2016;22(1):46–53.
- Badders NM, et al. Selective modulation of the androgen receptor AF2 domain rescues degeneration in spinal bulbar muscular atrophy. Nat Med. 2018;24(4):427–437.
- de Jong A, Schuurman KG, Rodenko B, Ovaa H, Berkers CR. Fluorescence-based proteasome activity profiling. Methods Mol Biol. 2012;803:183–204.
- Blount JR, et al. Ubiquitin-binding site 2 of ataxin-3 prevents its proteasomal degradation by interacting with Rad23. Nat Commun. 2014;5:4638. View this article via: PubMed Google Scholar
- Tsou WL, et al. DnaJ-1 and karyopherin α3 suppress degeneration in a new Drosophila model of spinocerebellar ataxia type 6. Hum Mol Genet. 2015;24(15):4385–4396.
- Tsou WL, et al. The deubiquitinase ataxin-3 requires Rad23 and DnaJ-1 for its neuroprotective role in Drosophila melanogaster. Neurobiol Dis. 2015;82:12–21.
-
Version history
- Version 1 (May 29, 2018): In-Press Preview
- Version 2 (July 23, 2018): Electronic publication
- Version 3 (August 1, 2018): Print issue publication



Copyright © 2024 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)
